| 核酸检测方法与流程 | 您所在的位置:网站首页 › c2c1基因 › 核酸检测方法与流程 |
核酸检测方法与流程
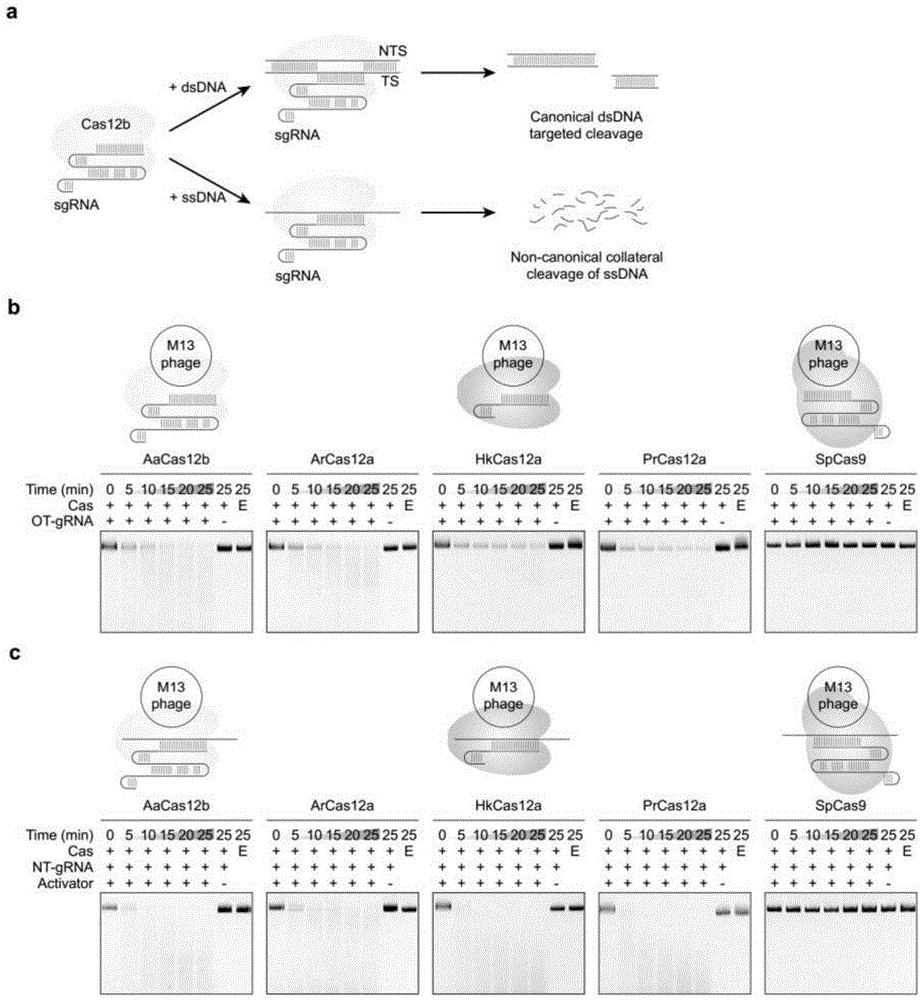 本发明涉及基因工程领域。具体而言,本发明涉及新的基于crispr系统的核酸检测方法。更具体而言,本发明涉及cas12b介导的dna检测方法及相关试剂盒。 背景技术: 快速便携的核酸检测有望在临床诊断和检疫检测中发挥重要作用。crispr核酸酶cas12a和cas13,由于具有ssdna或ssrna反式切割活性已经被开发用于以高灵敏度和特异性快速检测dna和rna。基于crispr-cas13的rna检测平台称为sherlock,基于cas12a的dna检测平台也称作detectr。 开发进一步的具有更高灵敏度和更高特异性的核酸检测平台在本领域具有重要意义。 发明简述 本发明提供一种基于cas12b的核酸检测方法,称为cdetection,能够以比cas12a更高的特异性检测dna,并且具有高达亚渺摩尔(attomolar)级的灵敏度。本发明还提供了增强的cdetection(ecdetection),其能够实现区分仅有单个核苷酸多态性不同的两种靶dna。基于crispr-cas12b的cdetection技术能够提供快速简便的dna检测方法,适用于健康领域和生物技术的各种应用。 附图描述 图1.cas12蛋白中保守的靶识别激活非特异性单链dna(ssdna)切割。(a)示意图说明cas12b具有经典的dsdna靶识别和切割,以及非经典的旁路ssdna切割。(b)m13mp18ssdna底物切割时间曲线,其中纯化的aacas12b、arcas12a、hkcas12a、prcas12a和spcas9和与m13噬菌体互补的向导rna(靶grna,ot(ontarget)-grna)结合。(c)m13mp18ssdna底物切割时间过程,其中纯化的aacas12b、arcas12a、hkcas12a、prcas12a和spcas9和与m13噬菌体没有序列同源性的grna(非靶grna,nt(non-target)-grna)及互补ssdna激活物结合。 图2.cas12b的ruvc结构域负责ssdna反式切割。(a-b)m13mp18ssdna底物和puc19dsdna切割时间曲线,其中纯化的wtaacas12b和ruvc催化突变体(r785a或d977a)与靶grna(ot-grna)或非靶grna(nt-grna)结合(a),或与nt-grna及互补ssdna激活物(b)结合,其中非靶grna与m13噬菌体或puc19没有序列同源性。 图3.cas12b介导的反式激活切割对非特异性ssdna的偏好性。(a)aacas12b介导的对含有a、t、g或c核苷酸同聚体的ssdna报告分子的反式切割的核苷酸偏好性。将aacas12b与靶向合成ssdna靶1或靶2的sgrna一起孵育。误差棒表示平均值的标准误差(s.e.m.),n=3。rfu,相对荧光单位。(b)测试多种缓冲液对cas12b介导的反式激活切割的影响。将aacas12b与靶向合成ssdna靶1的sgrna一起孵育。误差棒表示平均值的标准误差(s.e.m.),n=3。(c)使用on-target-ssdna(靶-ssdna或ot-ssdna)、non-target-ssdna(非靶-ssdna或nt-ssdna)、on-target-dsdna(靶-dsdna或ot-dsdna)和non-target-dsdna(非靶-dsdna或nt-dsdna)作为激活物,对aacas12b的反式切割活性分析。误差条表示s.e.m.,n=3。 图4.cas12b介导的dna检测的特异性和灵敏度。(a)使用具有指示的单个错配的ssdna或dsdna激活物分析aacas12b的反式切割活性。误差条表示s.e.m.,n=3。rfu,相对荧光单位。pt,完美配对的靶。mpam,突变的pam。(b)使用具有指示的连续错配的ssdna或dsdna激活物分析aacas12b的反式切割活性。误差棒表示s.e.m.,n=3。(c)比较aacas12b、prcas12a、lbcas12a在区分dsdna方面的特异性,使用合成的hpv16激活物。误差条表示s.e.m.,n=3。(d)使用dsdna激活物比较aac2c1的反式切割活性和预扩增增强的反式切割活性(cdetection)。将aacas12b与靶向靶1合成dsdna的sgrna一起温育。误差条表示s.e.m.,n=3。rpa,重组酶聚合酶扩增。(e)基于aacas12b、prcas12a、lbcas12a的使用rpa预扩增的dna检测所获得的最大荧光信号。将cas12蛋白与靶向合成hpv16dsdna的grna孵育,所述合成hpv16dsdna与背景基因组混合。误差条表示s.e.m.,n=3。(f)基于aacas12b、lbcas12a的使用rpa预扩增的dna检测所获得的荧光时间曲线。将cas12蛋白与靶向合成hpv16dsdna的grna孵育,所述合成hpv16dsdna稀释于人血浆中,终浓度10-18m。误差条表示s.e.m.,n=3。 图5.aacas12b介导的dna检测的特异性和灵敏度。(a)比较没有预扩增的使用ssdna或dsdna激活物的aacas12b反式切割活性。aacas12b与靶向合成ssdna或dsdna的sgrna一起孵育。误差条表示s.e.m.,n=3。(b)aacas12b检测到camvdna的存在。(c)aacas12b区分两种密切相关的合成hpv序列,其具有六个核苷酸多态性。误差棒表示s.e.m.,n=3。(d)比较aacas12b、prcas12a、lbcas12a在区分dsdna方面的特异性,使用合成的hpv18激活物。误差条表示s.e.m.,n=3。(e)使用dsdna激活物比较aacas12b的反式切割活性和预扩增增强的反式切割活性(cdetection)。将aacas12b与靶向靶2的合成ssdna或dsdna的sgrna一起温育。误差条表示s.e.m.,n=3。 图6.cdetection在dna检测中达到亚渺摩尔级灵敏度。(a)在检测和背景基因组混合的hpv16或hpv18dsdna时,cdetection实现亚渺摩尔级灵敏度。误差棒表示平均值的标准误差(s.e.m.),n=3。rfu,相对荧光单位。(b)使用rpa预扩增,从基于aacas12b和lbcas12a的dna检测中获得的荧光时间曲线。cas12蛋白与靶向合成hpv18dsdna的grna混合,所述合成hpv18dsdna稀释于人血浆中,终浓度10-18m。误差条表示s.e.m.,n=3。 图7.cdetection的应用。(a)通过cdetection检测abo血液基因分型的示意图。示出六种常见abo等位基因和三种靶向性sgrna。各sgrna以可检测信号区分出各自对应的等位基因。如果所有sgrna均没有产生信号,则等位基因为a101或a201。(b)使用cdetection进行abo血液基因分型获得的荧光信号。cdetection未能区分仅有一个单碱基多态性不同的两种dsdna激活物(on-b101vs.off-b101,on-o01vs.off-o01,on-o02/03vs.off-o02/03)。误差棒表示平均值的标准误差(s.e.m.),n=3。rfu,相对荧光单位。(c)示意通过引入经调节的grna(tgrna)开发ecdetection。使用sgrna的cdetection无法区分两种仅有单一错配的dsdna激活物,而使用tgrna(其在间隔区携带错配)的ecdetecion可以通过单核苷酸分辨率实现dna检测。 图8.增强型cdetection(ecdetection)的广泛应用。(a)使用tgrna的ecdection以单核苷酸分辨率特异性实现abo血液基因分型检测。误差棒表示s.e.m.,n=3。rfu,相对荧光单位。tgrna,经调节的grna。(b)和(c)上图示意brca1基因和靶向sgrna以及tgrna之间的序列差异。下图用最大荧光信号示出无rpa的cdetection使用sgrna和tgrna检测人brca1基因突变3232a>g(b)和3537a>g(c)的特异性。(d)荧光时间曲线显示有rpa的cdetection使用tgrna(3232-1)检测人brca1基因突变3232a>g的灵敏度和特异性。野生型brca1或brca13232a>gdsdna稀释于人血浆中,终浓度为10-18m。误差棒表示s.e.m.,n=3。 图9.使用cdetection平台进行精确dna检测。(a)上图示出tp53基因和靶向sgrna以及tgrna之间的序列差异。下图用最大荧光信号示出使用sgrna和tgrna检测人tp53基因突变856g>a的特异性。误差棒表示s.e.m.,n=3。rfu,相对荧光单位。tgrna,经调节的grna。(b)和(c)上图示意brca1基因和靶向sgrna以及tgrna之间的序列差异。下图用最大荧光信号示出使用sgrna和tgrna检测人brca1基因突变3232a>g(b)和3537a>g(c)的特异性。(d)荧光时间曲线显示有rpa的cdetection使用tgrna(3232-1)检测人brca1基因突变3232a>g的灵敏度和特异性。野生型brca1或brca13232a>gdsdna稀释于人血浆中,终浓度为10-16m。误差棒表示s.e.m.,n=3。 图10.cdetection平台的快速及精确的广泛诊断应用。首先通过直接裂解从不同样品获得基因组dna用于临床诊断和定量。对于不同的目的,经过或未经rpa预扩增的dna通过使用sgrna或tgrna的cdetection进行检测。 图11.选择用于基因组编辑测试的非冗余c2c1直系同源物的系统发生树及其基因座。(a)邻接系统发生树,显示测试的c2c1直系同源物的进化关系。(b)对应于(a)中突出显示的8种c2c1蛋白的细菌基因座图谱crrnadr和推定的tracrrna的模拟共折叠显示出稳定的二级结构。dr,直接重复。每个细菌基因组间隔区(spacer)的数目在其crispr阵列的上方或下方表示。 图12.c2c1直系同源物的蛋白质比对:测试的10种c2c1直系同源物的氨基酸序列的多序列比对。保守的残基用红色背景突出显示,保守突变用轮廓和红色字体突出显示。 图13.人293t细胞中c2c1直系同源物介导的基因组靶向。(a)t7ei测定结果表明在人类基因组中与其同源sgrna结合的八种c2c1蛋白的基因组靶向活性。三角形表示切割的条带。(b)t7ei测定结果表明在人293t细胞中由与其同源sgrna(bs3sgrna)结合的bs3c2c1介导的同时多重基因组靶向。(c)sanger测序显示由与bs3sgrna结合的bs3c2c1诱导的代表性插入缺失(indel)。pam和原间隔区序列分别用红色和蓝色着色。插入缺失和插入分别用紫色破折号和绿色小写字符表示。 图14.用于rna指导的基因组编辑的c2c1蛋白。(a)本发明中测试的10种c2c1直系同源物的图形概述。示出其大小(氨基酸数目)。(b)t7ei测定结果表明在人293t细胞中由其同源sgrna指导的八种c2c1直系同源物的基因组靶向活性。三角形表示切割的条带。(c-d)t7ei测定结果表明在人293t细胞中由aasgrna(c)和aksgrna(d)指导的八种c2c1直系同源物的基因组靶向活性。三角形表示切割的条带。 图15.c2c1的sgrna的dna比对:测试衍生自10个c2c1基因座的8种sgrna的dna序列的多序列比对。 图16.不同c2c1直系同源物与sgrna之间的可互换性。t7ei测定结果表明在人293t细胞中由aasgrna(a)、aksgrna(b)、amsgrna(c)、bs3sgrna(d)和lssgrna(e)指导的八种c2c1直系同源物的基因组靶向活性。红色三角形表示切割的条带。 图17.人工sgrna介导的多重基因组靶向。(a)对应于dic2c1和tcc2c1的细菌基因座的图谱。两个c2c1基因座没有crispr阵列。(b-c)t7ei测定结果表明在人293t细胞中由aasgrna(b)和aksgrna(c)指导的aac2c1、dic2c1和tcc2c1的基因组靶向活性。三角形表示切割的条带。(d)t7ei测定结果表明在人293t细胞中由与aksgrna结合的tcc2c1介导的同时多重基因组靶向。(e)示意图说明人工sgrna支架13(artgrna13)的二级结构。(f)t7ei测定结果表明在人293t细胞中由与artgrna13结合的tcc2c1介导的同时多重基因组靶向。 图18.不同sgrna指导c2c1进行基因组编辑。t7ei测定结果表明在人293t细胞中由aasgrna(a)、aksgrna(b)、amsgrna(c)、bs3sgrna(d)和lssgrna(e)指导的aac2c1、dic2c1和tcc2c1的基因组靶向活性。三角形表示切割的条带。 图19.tcc2c1介导的多重基因组编辑。(a)t7ei测定结果表明在人293t细胞中由与amsgrna结合的tcc2c1介导的同时多重基因组靶向。(b-c)sanger测序显示由与aksgrna(b)和amsgrna(c)结合的tcc2c1诱导的代表性插入缺失。pam和原间隔区序列分别用红色和蓝色着色。插入缺失和插入分别用紫色破折号和绿色小写字符表示。 图20.人工sgrna指导tcc2c1进行基因组编辑。(a)示意图说明36种人工sgrna(artgrna)支架(支架:1-12和14-37)的二级结构。(b)t7ei测定结果表明在人293t细胞中artsgrna指导的tcc2c1的基因组靶向活性。三角形表示切割的条带。(c)t7ei测定结果表明在人293t细胞中由与artgrna13结合的aac2c1介导的同时多重基因组靶向。 发明详述 在本发明中,除非另有说明,否则本文中使用的科学和技术名词具有本领域技术人员所通常理解的含义。并且,本文中所用的蛋白质和核酸化学、分子生物学、细胞和组织培养、微生物学、免疫学相关术语和实验室操作步骤均为相应领域内广泛使用的术语和常规步骤。例如,本发明中使用的标准重组dna和分子克隆技术为本领域技术人员熟知,并且在如下文献中有更全面的描述:sambrook,j.,fritsch,e.f.和maniatis,t.,molecularcloning:alaboratorymanual;coldspringharborlaboratorypress:coldspringharbor,1989(下文称为“sambrook”)。同时,为了更好地理解本发明,下面提供相关术语的定义和解释。 在第一方面,本发明提供一种检测生物样品中靶核酸分子的存在和/或量的方法,所述方法包括以下步骤: (a)使所述生物样品接触:i)cas12b蛋白,ii)针对所述靶核酸分子中的靶序列的grna,和iii)被切割后产生可检测信号的单链dna报告分子,从而形成反应混合物; (b)检测所述反应混合物中产生的可检测信号的存在和/或水平, 其中所述可检测信号的存在和/或水平代表所述靶核酸分子的存在和/或量。 在一些实施方案中,所述靶核酸分子是双链dna分子。在一些实施方案中,所述靶核酸分子是单链dna分子。所述靶核酸分子可以是基因组dna、cdna、病毒dna等,或它们的片段。 “cas12b”、“cas12b核酸酶”、“cas12b蛋白”、“c2c1”、“c2c1核酸酶”和“c2c1蛋白”在本文中可互换使用,指的是一种来自微生物crispr系统的rna指导的序列特异性核酸酶。cas12b能在向导rna的指导下靶向并切割dna靶序列形成dna双链断裂(dsb),也称作经典dsdna切割活性。更重要的是,cas12b与grna的复合物在识别并结合相应靶dna序列后,能够激活其非特异性单链dna切割活性,也称作非经典旁路ssdna切割活性。利用非经典旁路ssdna切割活性,切割在被切割后产生可检测信号的单链dna报告分子,即可以反映靶dna的存在和/或量。在本文中,被cas12b与grna的复合物识别并结合,激活cas12b的非特异性单链dna切割活性的dna分子也称作“激活物”。 在一些实施方案中,所述cas12b蛋白是来自alicyclobacillusacidiphilus的aacas12b蛋白、来自alicyclobacilluskakegawensis的akcas12b蛋白、来自alicyclobacillusmacrosporangiidus的amcas12b蛋白、来自bacillushisashii的bhcas12b蛋白、来自bacillus属的bscas12b蛋白、来自bacillus属的bs3cas12b蛋白、来自desulfovibrioinopinatus的dicas12b蛋白、来自laceyellasediminis的lscas12b蛋白、来自spirochaetesbacterium的sbcas12b蛋白、来自tuberibacilluscalidus的tccas12b蛋白。在一些优选实施方案中,所述cas12b蛋白是来自alicyclobacillusacidiphilus的cas12b蛋白(aacas12b)。本申请人已经鉴定这些cas12b蛋白可以用于在哺乳动物中进行基因组编辑,也可以用于本发明的核酸检测方法。 例如,所述cas12b蛋白是来自alicyclobacillusacidiphilusnbrc100859的aacas12b蛋白、来自alicyclobacilluskakegawensisnbrc103104的akcas12b蛋白、来自alicyclobacillusmacrosporangiidusstraindsm17980的amcas12b蛋白、来自bacillushisashiistrainc4的bhcas12b蛋白、来自bacillus属nsp2.1的bscas12b蛋白、来自bacillus属v3-13contig_40的bs3cas12b蛋白、来自desulfovibrioinopinatusdsm10711的dicas12b蛋白、来自laceyellasediminisstrainrha1的lscas12b蛋白、来自spirochaetesbacteriumgwb1_27_13的sbcas12b蛋白、来自tuberibacilluscalidusdsm17572的tccas12b蛋白。在一些优选实施方案中,所述cas12b蛋白是来自alicyclobacillusacidiphilusnbrc100859的cas12b蛋白。 alicyclobacillusacidiphilus的cas12b的基因座上缺少已经测序的直接重复(dr)阵列,因此本领域技术人员将会认为其无法进行基因编辑,从而在crispr核酸酶筛选中将其略过。然而,本发明人令人惊奇地发现,来自alicyclobacillusacidiphilus的cas12b蛋白同样具有经典靶向dsdna切割活性和非经典旁路ssdna切割活性,从而能用于基因编辑和核酸检测。类似地,所鉴定的其它一些cas12b蛋白,例如dicas12b或tccas12b蛋白,尽管其天然基因座不具有crispr阵列,也出乎意料地可以用于本发明。 在本发明一些实施方式中,所述cas12b蛋白是其天然基因座不具有crispr阵列的cas12b蛋白。在一些实施方式中,所述天然基因座不具有crispr阵列的cas12b蛋白是aacas12b蛋白、dicas12b或tccas12b蛋白。 在一些实施方案中,所述cas12b蛋白包含与seqidno:1-10中任一个具有至少80%、至少85%、至少90%、至少95%、至少96%、至少97%、至少98%、至少99%序列相同性的氨基酸序列。在一些实施方案中,所述cas12b蛋白的氨基酸序列相对于seqidno:1-10中任一个具有一或多个氨基酸残基取代、缺失或添加。例如,所述cas12b蛋白相对于seqidno:1-10中任一个具有1个、2个、3个、4个、5个、6个、7个、8个、9个或10个氨基酸残基取代、缺失或添加的氨基酸序列。在一些实施方案中,所述氨基酸取代是保守型取代。在一些实施方案中,所述cas12b蛋白包含seqidno:1-10中任一所示的氨基酸序列。例如,所述aacas12b、akcas12b、amcas12b、bhcas12b、bscas12b、bs3cas12b、dicas12b、lscas12b、sbcas12b、tccas12b蛋白分别包含seqidno:1-10所示氨基酸序列。在一些优选实施方式中,所述cas12b蛋白包含seqidno:1所示的氨基酸序列。 本发明人已经证明了cas12b蛋白的ruvc结构域对于其非经典旁路ssdna切割活性是关键的。在一些实施方案中,所述cas12b蛋白包含野生型cas12b蛋白的ruvc结构域,所述野生型cas12b蛋白例如包含seqidno:1-10任一项所示氨基酸序列。本领域技术人员可以容易地鉴定出cas12b蛋白的ruvc结构域。例如,通过ncbi提供的工具。 序列“相同性”具有本领域公认的含义,并且可以利用公开的技术计算两个核酸或多肽分子或区域之间序列相同性的百分比。可以沿着多核苷酸或多肽的全长或者沿着该分子的区域测量序列相同性。(参见,例如:computationalmolecularbiology,lesk,a.m.,ed.,oxforduniversitypress,newyork,1988;biocomputing:informaticsandgenomeprojects,smith,d.w.,ed.,academicpress,newyork,1993;computeranalysisofsequencedata,parti,griffin,a.m.,andgriffin,h.g.,eds.,humanapress,newjersey,1994;sequenceanalysisinmolecularbiology,vonheinje,g.,academicpress,1987;andsequenceanalysisprimer,gribskov,m.anddevereux,j.,eds.,mstocktonpress,newyork,1991)。虽然存在许多测量两个多核苷酸或多肽之间的相同性的方法,但是术语“相同性”是技术人员公知的(carrillo,h.&lipman,d.,siamjappliedmath48:1073(1988))。 在肽或蛋白中,合适的保守型氨基酸取代是本领域技术人员已知的,并且一般可以进行而不改变所得分子的生物活性。通常,本领域技术人员认识到多肽的非必需区中的单个氨基酸取代基本上不改变生物活性(参见,例如,watsonetal.,molecularbiologyofthegene,4thedition,1987,thebenjamin/cummingspub.co.,p.224)。 特别地,本领域人员将可以理解,在同一细菌物种的不同菌株cas12b蛋白可能在氨基酸序列存在一定差异,但是却能实现基本上相同的功能。 在一些实施方案中,所述cas12b蛋白是重组产生的。在一些实施方案中,所述cas12b蛋白还含有融合标签,例如用于cas12b蛋白分离/和或纯化的标签。重组产生蛋白质的方法是本领域已知的。并且本领域已知多种可以用于分离/和或纯化蛋白质的标签,包括但不限于his标签、gst标签等。通常而言,这些标签不会改变目的蛋白的活性。 向导rna”和“grna”在本文中可互换使用,通常由部分互补形成复合物的crrna和tracrrna分子构成,其中crrna包含与靶序列具有足够相同性以便与靶序列的互补序列杂交并且指导crispr复合物(crispr核酸酶+crrna+tracrrna)与该靶序列以序列特异性方式结合的序列。然而,可以设计并使用单向导rna(sgrna),其同时包含crrna和tracrrna的特征。不同的crispr核酸酶对应的grna存在不同。例如,cas9和cas12b通常需要crrna和tracrrna两者,然而,cas12a(cpf1)则只需要crrna。 “针对靶核酸分子的靶序列的grna”指的是grna能够特异性识别所述靶序列。例如,在一些实施方案中(靶核酸分子是双链dna),所述grna包含能够与靶序列的互补序列特异性杂交的间隔区序列(spacer)。在一些实施方案中(靶核酸分子是单链dna),所述grna包含能够与靶序列特异性杂交的间隔区序列。 a.acidiphiluscrispr基因座中没有直接重复序列(dr)阵列,因此,aacas12b并没有对应的crrna。然而,本发明人发现,aacas12b也可以采用来自其他生物体的cas12b蛋白的相应grna。例如aacas12b可以采用自身的tracrrna和来自a.acidoterrestris的crispr基因座的crrna序列作为grna。本发明人对可用于aacas12b的grna进行了优化。 在本发明的一些实施方案中,所述向导rna是由crrna和tracrrna部分互补形成的复合物。在一些实施方案中,所述tracrrna由以下的核酸序列编码:5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgctcgctcagtgttctgac-3’。在一些实施方案中,所述crrna由以下的核酸序列编码:5’-gtcggatcactgagcgagcgatctgagaagtggcac-nx-3’,其中nx表示x个连续的核苷酸组成的核苷酸序列,n各自独立地选自a、g、c和t;x为18≤x≤35的整数。优选地,x=20。在一些实施方案中,nx是能够与靶序列的互补序列特异性杂交的间隔区序列(靶核酸分子是双链dna)。在一些实施方案中,nx是能够与靶序列特异性杂交的间隔区序列(靶核酸分子是单链dna)。 在本发明的一些实施方案中,所述向导rna是sgrna。在一些实施方案中,所述sgrna由5’端的支架序列和3’端的间隔区序列组成。间隔区序列可以与靶序列或靶序列的互补序列特异性杂交。间隔区序列通常长度为18至35个核苷酸,优选20个核苷酸。 在一些具体实施方案中,所述sgrna由选自以下之一的核酸序列编码: 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcgagcgatctgagaagtggcac-nx-3’; 5’-aactgtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcgagcgatctgagaagtggcac-nx-3’; 5’-ctgtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcgagcgatctgagaagtggcac-nx-3’; 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgctcgctcagtgttatcactgagcgagcgatctgagaagtggcac-nx-3’; 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgatctgagaagtggcac-nx-3’; 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagctgagaagtggcac-nx-3’; 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaagctgagaagtggcac-nx-3’; 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaactgagaagtggcac-nx-3’; 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaagcgagaagtggcac-nx-3’; 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctaagcagaagtggcac-nx-3’;和 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’; 其中nx表示x个连续的核苷酸组成的核苷酸序列,n各自独立地选自a、g、c和t;x为18≤x≤35的整数。优选地,x=20。在一些实施方案中,nx是能够与靶序列的互补序列特异性杂交的间隔区序列(靶核酸分子是双链dna)。在一些实施方案中,nx是能够与靶序列特异性杂交的间隔区序列(靶核酸分子是单链dna)。在一些实施方案中,所述sgrna包含由seqidno:11-21中任一项的核苷酸序列编码的支架序列。 在一些具体实施方案中,所述sgrna由选自以下之一的核酸序列编码: 5’-gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcgagcgatctgagaagtggcac-nx-3’(aasgrna); 5’-tcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaagatgaccgctcgctcagcgatctgacaacggatcgctgagcgagcggtctgagaagtggcac-nx-3’(aksgrna1); 5’-ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaacatgatcgcccgctcaacggtccgatgtcggatcgttgagcgggcgatctgagaagtggcac-nx-3’(amsgrna1); 5’-gaggttctgtcttttggtcaggacaaccgtctagctataagtgctgcagggtgtgagaaactcctattgctggacgatgtctcttttatttcttttttcttggatgtccaagaaaaaagaaatgatacgaggcattagcac-nx-3’(bhsgrna); 5’-ccataagtcgacttacatatccgtgcgtgtgcattatgggcccatccacaggtctattcccacggataatcacgactttccactaagctttcgaatgttcgaaagcttagtggaaagcttcgtggttagcac-nx-3’(bssgrna); 5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatcttatttctgctaagtgtttagttgcctgaatacttagcagaaataatgatgattggcac-nx-3’(bs3sgrna); 5’-ggcaaagaatactgtgcgtgtgctaaggatggaaaaaatccattcaaccacaggattacattatttatctaatcacttaaatctttaagtgattagatgaattaaatgtgattagcac-nx-3’(lssgrna);或 5’-gtcttagggtatatcccaaatttgtcttagtatgtgcattgcttacagcgacaactaaggtttgtttatcttttttttacattgtaagatgttttacattataaaaagaagataatcttattgcac-nx-3’(sbsgrna); 其中nx表示x个连续的核苷酸组成的核苷酸序列(spacer序列),n各自独立地选自a、g、c和t;x为18≤x≤35的整数。优选地,x=20。在一些实施方案中,序列nx(spacer序列)能够与靶序列的互补序列特异性杂交。所述sgrna中除nx之外的序列为sgrna的支架(scaffold)序列。在一些实施方案中,所述sgrna包含由seqidno:22-29中任一项的核苷酸序列编码的支架序列。 本发明令人惊奇地发现,不同的cas12b系统中的cas12b蛋白以及向导rna可以互换使用,从而使得可以人工设计通用的向导rna。 因此在一些实施方案中,所述sgrna是人工sgrna,其由选自以下的核苷酸序列编码: 5’-ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna1); 5’-ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna2); 5’-ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacagacttcaagcgaagtggcac-nx-3’(artsgrna3); 5’-ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna4); 5’-ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna5); 5’-ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccaccacagacttcaagcgaagtggcac-nx-3’(artsgrna6); 5’-ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna7); 5’-ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna8); 5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacagacttcaagcgaagtggcac-nx-3’(artsgrna9); 5’-ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna10); 5’-ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna11); 5’-ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggcttcaaagaagtggcac-nx-3’(artsgrna12); 5’-ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna13); 5’-ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna14); 5’-ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggcttcaaagaagtggcac-nx-3’(artsgrna15); 5’-ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna16); 5’-ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna17); 5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggcttcaaagaagtggcac-nx-3’(artsgrna18); 5’-ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagattatctatgatgattggcac-nx-3’(artsgrna19); 5’-ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggattatctatgatgattggcac-nx-3’(artsgrna20); 5’-ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatctatgatgattggcac-nx-3’(artsgrna21); 5’-ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagattatctatgatgattggcac-nx-3’(artsgrna22); 5’-ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggattatctatgatgattggcac-nx-3’(artsgrna23); 5’-ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatctatgatgattggcac-nx-3’(artsgrna24); 5’-ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagattatctatgatgattggcac-nx-3’(artsgrna25); 5’-ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggattatctatgatgattggcac-nx-3’(artsgrna26); 5’-ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccaccacaggattatctatgatgattggcac-nx-3’(artsgrna27); 5’-ggtctaaaggacagaacaacgggatgtgccaatgcactctttccaggagtgaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna28); 5’-ggtcgtctataggacggcgagcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna29); 5’-ggaattgccgatctataggacggcagatttttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgaacttcaagcgaagtggcac-nx-3’(artsgrna30); 5’-ggaattgccgatctataggacggcagattgacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna31); 5’-ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttgacttcaagcgaagtggcac-nx-3’(artsgrna32); 5’-ggtctaaaggacagaacaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna33); 5’-ggtcgtctataggacggcgagcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna34); 5’-ggaattgccgatctataggacggcagatttttttcaacgggtgtgccaatggccactttccaggtggcaaagcccgttgagcttcaaagaagtggcac-nx-3’(artsgrna35); 5’-ggaattgccgatctataggacggcagattgacaacgggaagtgccaatgtgctctttccaagagcaaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrn36a);或 5’-ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccaggagtgaacaccccgttggcttcaaagaagtggcac-nx-3’(artsgrna37), 其中nx表示x个连续的核苷酸组成的核苷酸序列(spacer序列),n各自独立地选自a、g、c和t;x为18≤x≤35的整数。优选地,x=20。在一些实施方案中,序列nx(spacer序列)能够与靶序列的互补序列特异性杂交。所述sgrna中除nx之外的序列为sgrna的支架(scaffold)序列。 在一些实施方案中,所述人工sgrna包含由seqidno:30-66中任一项的核苷酸序列编码的支架序列。 在一些实施方案中,所述grna的间隔区序列被设计为与靶序列或其互补序列完全匹配。在一些实施方案中,所述grna的间隔区序列被设计为与靶序列或其互补序列具有至少一个核苷酸错配,例如具有一个核苷酸错配。这样的grna也称为经调节的(tuned)grna,所述核苷酸错配也成为调节位点。利用cas12b对sgrna与靶之间不同错配的耐受能力差异,设计为与靶序列或其互补序列具有核苷酸错配的grna能够区分靶序列中的单核苷酸多态性变异。在一些实施方案中,所述至少一个核苷酸错配的位置不同于所述单核苷酸多态性变异的位置。例如,经调节的sgrna与靶序列1在位置1具有一个核苷酸错配,而靶序列1与靶序列2在位置2具有单核苷酸多肽性,也即经调节的sgrna与靶序列2之间具有两个核苷酸错配。由于cas12b对错配数目的耐受性差异,其仅在靶序列1存在时产生可检测信号(只有1个错配),而靶序列2没有可检测信号(由于存在两个错配),从而可以将包含单核苷酸多态性的靶序列1和靶序列2区分开。本领域技术人员可以根据具体的靶序列筛选合适的调节位点。 本发明中,grna中除间隔区序列之外的序列也称做grna支架(scaffold)。 在一些实施方案中,所述grna体外转录产生。在一些实施方案中,所述grna通过化学合成产生。 在本发明的一些实施方案中,所述靶核酸分子包含特征性的长度为18-35个核苷酸,优选20个核苷酸的靶序列。在本发明的一些实施方案中,特别是涉及双链dna检测,所述靶序列在紧接其5’端具有选自:5’tttn-3’、5’attn-3’、5’gttn-3’、5’cttn-3’、5’ttc-3’、5’ttg-3’、5’tta-3’、5’ttt-3’、5’tan-3’、5’tgn-3’、5’tcn-3’和5’atc-3’的pam(前间区邻近基序)序列,优选5’tttn-3’。 “被切割后产生可检测信号的单链dna报告分子”例如可以在单链dna的两端分别包含荧光团和其猝灭基团。当单链dna未被切割时,由于猝灭基团存在,荧光团不发出荧光。而当cas12b-grna复合物被靶核酸分子激活,通过其非经典旁路ssdna切割活性切割所述单链dna报告分子的dna单链时,荧光团被释放,从而发出荧光。合适的荧光团及其相应猝灭基团,以及其标记核酸分子的方法在本领域是已知的。合适的荧光团包括但不限于fam、tex、hex、cy3或cy5。合适的猝灭基团包括但不限于bhq1、bhq2、bhq3或tamra。合适的荧光团-猝灭基团对包括但不限于fam-bhq1、tex-bhq2、cy5-bhq3、cy3-bhq1或fam-tamra。因此,在一些实施方案中,所述可检测信号是荧光信号。在一些实施方式中,所述荧光团是fam,所述猝灭基团是bhq1。 所述单链dna报告分子中单链dna的长度可以是大约2-100个核苷酸,例如2-5个、2-10个、2-15个、2-20个、2-25个、2-30个、2-40个或2至更多个核苷酸。所述单链dna报告分子中单链dna可以包含任意序列,但在一些实施方案中,polyg(聚鸟苷酸)除外。在一些实施方案中,所述单链dna报告分子中的单链dna可以选自polya(聚腺苷酸)、polyc(聚胞苷酸)或polyt(聚胸苷酸)。 在一些具体实施方式中,所述单链dna报告分子选自5’-fam-aaaaa-bhq1-3’、5’-fam-ttttt-bhq1-3’、和5’-fam-ccccc-bhq1-3’。 在本发明的方法的一些实施方案中,还包括在步骤(a)之前对所述生物样品中的核酸分子进行扩增的步骤。所述扩增包括但不限于pcr扩增或重组酶聚合酶扩增(recombinasepolymeraseamplification,rpa)。优选地,所述扩增是重组酶聚合酶扩增。 在一些实施方案中,所述重组酶聚合酶扩增进行约10分钟-约60分钟 在一些实施方案中,在与所述生物样品接触之前,所述cas12b蛋白已经与所述grna预先复合形成cas12b-grna复合物。 在一些实施方案中,所述步骤(a)的反应进行约20分钟-约180分钟,例如约20分钟、约30分钟、约40分钟、约50分钟、约60分钟、约90分钟、约120分钟,或之间的任何时间。 在一些实施方案中,步骤(a)在合适的缓冲液中进行。例如,所述缓冲液是nebuffertm2、nebuffertm2.1或buffer。在一些实施方式中,所述缓冲液包含终浓度50mmnacl、10mmtris-hcl、10mmmgcl2、1mmdtt,ph7.9。在一些实施方式中,所述缓冲液包含终浓度50mmnacl、10mmtris-hcl、10mmmgcl2、100μg/mlbsa,ph7.9。在一些实施方式中,所述缓冲液包含终浓度50mm乙酸钾,20mmtris-乙酸,10mm乙酸镁,100μg/mlbsa,ph7.9。 可用于本发明的方法的生物样品包括但不限于全血、血浆、血清、脑脊液、尿液、粪便、细胞或组织提取物等。所述生物样品涵盖提取自细胞或组织的核酸样品。 本发明的范围内还包括用于本发明的方法的试剂盒,该试剂盒包括用于实施本发明的方法的试剂,以及使用说明。例如,所述试剂盒可以包括cas12b蛋白(例如本发明的cas12b蛋白)、grna(例如包含本发明的grna支架)或用于产生grna(例如其包含本发明的grna支架)的试剂、单链dna报告分子(例如本发明的单链dna报告分子),合适的缓冲液、和/或核酸扩增试剂。试剂盒一般包括表明试剂盒内容物的预期用途和/或使用方法的标签。术语标签包括在试剂盒上或与试剂盒一起提供的或以其他方式随试剂盒提供的任何书面的或记录的材料。 本发明还提供了上文所定义的cas12b蛋白和/或包含本发明的支架的grna和/或用于产生包含本发明的支架的grna的试剂在制备用于本发明的方法的试剂盒中的用途。 实施例 实验材料和方法 蛋白表达和纯化 spcas9和lbcas12a蛋白商购获得(neb)。根据之前的报道纯化aacas12b、arcas12a、hkcas12a和prcas12a蛋白。简言之,bpk2014-cas12-his10蛋白在大肠杆菌菌株bl21(λde3)中表达,用0.5mmiptg在16℃诱导表达16小时。收获细胞沉淀并裂解,使用his60nisuperflowresin(takara)纯化。将纯化的cas12蛋白透析,浓缩,最后使用bca蛋白质测定试剂盒(thermofisher)定量。 核酸制备 dna寡核苷酸商购获得(genscript)。双链dna激活物通过pcr扩增获得并使用oligoclean&concentratorkit(zymoresearch)纯化。向导rna使用hiscribetmt7highyieldrnasynthesiskit(neb)体外转录并用microeluternacleanupkit(omega)纯化。 在指定反应中使用的背景基因组dna使用mousedirectpcrkit(bimake)纯化自人胚肾293t细胞。为了模拟无细胞dna(cfdna),dsdna以指定浓度稀释至人血浆中(thermofisher)。 荧光团猝灭剂(fq)-标记的报告测定 用30nmcas12、36nmgrna、混合在40ng背景基因组dna中(在指定的反应中)的40nm激活物(除非另有说明)、200nm定制合成的均聚物ssdnafq报告基因(表1)和nebuffertm2(除非另有说明)在corning384-孔聚苯乙烯nbs微孔板中进行20μl反应检测测定。将反应物在37℃下孵育以在荧光板读数器(bioteksynergy4)中指示时间,每5分钟测量荧光动力学(λex=485nm;λem=528nm,透射增益=61)。通过sigmaplot软件分析荧光结果。 重组酶聚合酶扩增(rpa)反应 根据制造商的方案,使用twistampbasic(twistdx)进行重组酶聚合酶扩增(rpa)反应。将含有不同dna输入量的50μlrpa反应系统在37℃下孵育10分钟。如上所述,将16μlrpa产物直接转移至20μl检测试验。 实施例1、cas12b的反式ssdna切割活性的表征 源自alicyclobacillusacidiphilusnbrc100859的crispr-cas12b核酸酶(aacas12b,氨基酸序列示于seqidno:1)已被用于哺乳动物基因组编辑,因为其具有经典的dsdna靶向切割能力(图1a)。为了表征cas12b的非经典反式旁路切割活性,使用cas12b、cas12a和cas9分别与其相应的向导rna(grna)组合进行体外ssdna切割测定。结果表明,aacas12b和cas12a(arcas12a、hkcas12a和prcas12a)在靶向m13的sgrna(ot-grna)存在下可诱导单链m13dna噬菌体快速降解,而spcas9则不能(图1b)。同样,aacas12b和cas12a在存在与m13噬菌体基因组没有序列同源性的非靶grna及其互补ssdna“激活物”的情况下也实现了m13降解(图1c)。催化失活的变体(发生r785a和d977a取代)中非经典旁路ssdna切割活性被消除,表明该旁路切割活性是ruvc结构域依赖性的(图2)。这些结果表明aacas12b-sgrna复合物一旦被与向导序列互补的dna触发,就可以获得非特异性ssdna反式切割活性。 实施例2、开发cas12b介导的dna检测系统 为了开发cas12b介导的dna检测系统,首先分析了aacas12b-sgrna复合物对荧光团-猝灭剂(fq)标记的同聚物报告分子的切割偏好。发现aacas12b偏好多聚胸苷酸(ployt)以及多聚腺苷酸(polya)和多聚胞苷酸(polyc),而多聚鸟苷酸(polyg)基本上不被切割(图3a)。同时,可以优化切割效率,其在nebuffertm2表现最佳(图3b)。然后,使用nebuffertm2中的polyt报告分子进行aacas12b介导的切割测定。使用sgrna-互补的ot-ssdna和ot-dsdna(ot表示ontarget,在靶),或sgrna-非互补的nt-ssdna和nt-dsdna(nt表述nontarget,非靶)作为激活物,发现了ot-ssdna和ot-dsdna激活物能够触发aacas12b切割fq-报告分子,尽管ot-dsdna效率较低(图3c)。 接下来使用带有不同错配的ssdna或dsdna激活物测试了反式切割活化的特异性。发现pam序列对于dsdna激活物触发的aacas12b的反式切割活性至关重要,并且对于ssdna不是必须的(图4a、b)。同时,还发现dsdna激活物和sgrna之间的错配会阻碍甚至消除反式切割活性,而ssdna激活物则对错配更耐受(图4a、b)。 然后确定了aacas12b-sgrna-激活物系统的灵敏度,发现没有预扩增,aacas12b在ssdna-激活物输入浓度g和3537a>g)。用选择的tgrna(tgrna-3232-1和tgrna-3537-4)进行cdetection可以很好地区分点突变,而sgrna几乎不支持点突变检测(图8b、c和图9b、c)。 此外,为了模拟通过cdetection使用cfdna早期临床检测原发疾病,将brca13232a>gdsdna稀释到人血浆中。研究结果表明,cdetection可以在单碱基分辨率下实现点突变检测(图8d和图9d)。本发明的ecdetection能够以单碱基分辨率在临床研究实现快速dna检测。 总之,本发明提供了cdetection平台,该平台基于cas12b核酸酶的非经典旁路ssdna切割特性,可以渺摩尔级灵敏度检测dna。同时,结合经调节的grna,开发了增强版(ecdetection),实现单核苷酸分辨率。本发明的cdetection和ecdetection平台可以在各种分子诊断应用中更容易地检测核酸,以及在临床研究中进行基因型分析(图10)。 实施例4、其他cas12b蛋白的鉴定 选择并从头合成来自不同细菌的六种代表性cas12b蛋白,以及之前报道的四种cas12b直系同源物,在人胚胎肾293t细胞中进行基因组编辑(图11、12和seqidno:1-10)。在这10种cas12b直系同源物中,来自d.inopinatus(dicas12b)和t.calidus(tccas12b)的cas12b既没有可预测的前体crisprrna(pre-crrna)也没有反式激活crrna(tracrrna)(图11b),提示这两种cas12b蛋白可能不适合基因组编辑应用。 为了进行哺乳动物基因组编辑,用单独的cas12b酶和其靶向含有适当pam的人内源基因座的同源单向导rna(sgrna)共转染293t细胞(图11)。t7核酸内切酶(t7ei)测定的结果显示,aacas12b、akcas12b、amcas12b、bhcas12b、bs3cas12b和lscas12b稳健地编辑人类基因组,尽管它们的靶向效率在不同的直系同源物之间和在不同的靶向位点不同(图11b和图13a)。还通过简单地使用多个sgrna,使用bs3cas12b实现多重基因组编辑,同时编辑人类基因组中的四个位点(图13b,c)。这些新发现的有功能的cas12b直系同源物扩展对基于cas12b的基因组编辑的选择,同时扩展了基于cas12b的dna检测的选择。 实施例5、不同cas12b及双rna的可互换性 为了研究cas12b系统中双rna(crrna和tracrrna)和蛋白质组分之间的可互换性,首先分析cas12b蛋白和双rna两者的保守性。除了cas12b直系同源物的保守氨基酸序列外(图14a和图12),前体crrna:tracrrna双链体的dna序列及其二级结构也表现出高保守性(图11b和图15)。接下来,用分别与来自8个cas12b系统的各sgrna复合的8种cas12b直系同源物,在293t细胞中进行基因组编辑。如t7ei测定的结果所示,衍生自aacas12b、akcas12b、amcas12b、bs3cas12b和lscas12b基因座的sgrna可以替代原始sgrna用于哺乳动物基因组编辑,尽管在不同cas12b直系同源物和sgrna之间的活性有所不同(图14c,d和图16)。这些结果证明不同cas12b和来自不同cas12b基因座的双rna之间的可互换性。 实施例6、利用天然基因座无crispr阵列的cas12b进行基因组编辑 本发明进一步选择两个基因座没有携带crispr阵列的cas12b直系同源物dicas12b和tccas12b进行后续实验(图17a)。基因座没有携带crispr阵列使得它们的crrna:tracrrna双链体的序列不可预测。在293t细胞中共转染与靶向不同基因组位点的衍生自其他8种cas12b直系同源物的基因座的sgrna组合的dicas12b和tccas12b以及aacas12b。t7ei测定结果表明衍生自aacas12b、akcas12b、amcas12b、bs3cas12b和lscas12b的sgrna使tccas12b能够稳健地编辑人类基因组(图17b、c和图18)。此外,aasgrna或aksgrna能够使tccas12b实现多重基因组编辑(图17d和图19)。上述结果表明在来自不同系统的cas12b和双链rna之间可互换性使得可能利用天然基因座不具有crispr阵列的cas12b直系同源物来编辑哺乳动物基因组。 实施例7、设计用于cas12b介导的基因组编辑的人工sgrna 不同cas12b系统中cas12b蛋白和双rna之间的可互换性有利于设计新的人工sgrna(artsgrna)支架以促进cas12b介导的基因组编辑。考虑到cas12b直系同源物中dna序列和二级结构的保守性(图11b和13),设计并从头合成37种sgrna支架(seqidno:30-66),用于靶向人ccr5基因座(图17e,图20a)。t7ei测定的结果表明22种artsgrna支架有效地工作(图10b)。为了验证artgrna的普遍适用性,使用artsgrna13指导tccas12b或aacas12b进行多重基因组编辑(图20a)。t7ei测定结果表明,artsgrna13同时促进tccas12b和aacas12b两者的多重基因组编辑(图17f和图20c)。结果表明通过设计和合成artsgrna能促进cas12b介导的基因组编辑特别是多重基因组编辑。 表1.实施例中涉及的核酸序列(其中下划线为spacer序列,粗体斜体为tgrna中的错配核苷酸) 表2.cas12b-介导的dna检测中测试的缓冲液的组分。缓冲液1、3、4、5来自已有的报道,而缓冲液2、6、7、8来自商品化的缓冲液。1:nucleaseassaybuffer;2:nebuffertm3.1;3:cas12abindingbuffer;4:cas13buffer;5:cas12abuffer;6:nebuffertm2;7:nebuffertm2.1;8:buffer。 表3.本发明的序列表及对应的序列信息 序列表 中国科学院动物研究所 核酸检测方法 tc2737 201811099146.0 2018-09-20 66 patentinversion3.5 1 1129 prt alicyclobacillusacidiphilus 1 metalavallyssermetlysvallysleuargleuaspasnmetpro 151015 gluileargalaglyleutrplysleuhisthrgluvalasnalagly 202530 valargtyrtyrthrglutrpleuserleuleuargglngluasnleu 354045 tyrargargserproasnglyaspglygluglnglucystyrlysthr 505560 alagluglucyslysalagluleuleugluargleuargalaarggln 65707580 valgluasnglyhiscysglyproalaglyseraspaspgluleuleu 859095 glnleualaargglnleutyrgluleuleuvalproglnalailegly 100105110 alalysglyaspalaglnglnilealaarglyspheleuserproleu 115120125 alaasplysaspalavalglyglyleuglyilealalysalaglyasn 130135140 lysproargtrpvalargmetargglualaglygluproglytrpglu 145150155160 gluglulysalalysalaglualaarglysserthraspargthrala 165170175 aspvalleuargalaleualaasppheglyleulysproleumetarg 180185190 valtyrthraspseraspmetserservalglntrplysproleuarg 195200205 lysglyglnalavalargthrtrpaspargaspmetpheglnglnala 210215220 ilegluargmetmetsertrpglusertrpasnglnargvalglyglu 225230235240 alatyralalysleuvalgluglnlysserargphegluglnlysasn 245250255 phevalglyglngluhisleuvalglnleuvalasnglnleuglngln 260265270 aspmetlysglualaserhisglyleugluserlysgluglnthrala 275280285 histyrleuthrglyargalaleuargglyserasplysvalpheglu 290295300 lystrpglulysleuaspproaspalapropheaspleutyraspthr 305310315320 gluilelysasnvalglnargargasnthrargargpheglyserhis 325330335 aspleuphealalysleualagluprolystyrglnalaleutrparg 340345350 gluaspalaserpheleuthrargtyralavaltyrasnserileval 355360365 arglysleuasnhisalalysmetphealathrphethrleuproasp 370375380 alathralahisproiletrpthrargpheasplysleuglyglyasn 385390395400 leuhisglntyrthrpheleupheasnglupheglygluglyarghis 405410415 alaileargpheglnlysleuleuthrvalgluaspglyvalalalys 420425430 gluvalaspaspvalthrvalproilesermetseralaglnleuasp 435440445 aspleuleuproargaspprohisgluleuvalalaleutyrphegln 450455460 asptyrglyalagluglnhisleualaglyglupheglyglyalalys 465470475480 ileglntyrargargaspglnleuasnhisleuhisalaargarggly 485490495 alaargaspvaltyrleuasnleuservalargvalglnserglnser 500505510 glualaargglygluargargproprotyralaalavalpheargleu 515520525 valglyaspasnhisargalaphevalhispheasplysleuserasp 530535540 tyrleualagluhisproaspaspglylysleuglysergluglyleu 545550555560 leuserglyleuargvalmetservalaspleuglyleuargthrser 565570575 alaserileservalpheargvalalaarglysaspgluleulyspro 580585590 asnsergluglyargvalprophecyspheproilegluglyasnglu 595600605 asnleuvalalavalhisgluargserglnleuleulysleuprogly 610615620 gluthrgluserlysaspleuargalaileargglugluargglnarg 625630635640 thrleuargglnleuargthrglnleualatyrleuargleuleuval 645650655 argcysglysergluaspvalglyargarggluargsertrpalalys 660665670 leuilegluglnprometaspalaasnglnmetthrproasptrparg 675680685 glualaphegluaspgluleuglnlysleulysserleutyrglyile 690695700 cysglyaspargglutrpthrglualavaltyrgluservalargarg 705710715720 valtrparghismetglylysglnvalargasptrparglysaspval 725730735 argserglygluargprolysileargglytyrglnlysaspvalval 740745750 glyglyasnserilegluglnileglutyrleugluargglntyrlys 755760765 pheleulyssertrpserphepheglylysvalserglyglnvalile 770775780 argalaglulysglyserargphealailethrleuarggluhisile 785790795800 asphisalalysgluaspargleulyslysleualaaspargileile 805810815 metglualaleuglytyrvaltyralaleuaspaspgluargglylys 820825830 glylystrpvalalalystyrproprocysglnleuileleuleuglu 835840845 gluleuserglutyrglnpheasnasnaspargproprosergluasn 850855860 asnglnleumetglntrpserhisargglyvalpheglngluleuleu 865870875880 asnglnalaglnvalhisaspleuleuvalglythrmettyralaala 885890895 pheserserargpheaspalaargthrglyalaproglyileargcys 900905910 argargvalproalaargcysalaarggluglnasnprogluprophe 915920925 protrptrpleuasnlysphevalalagluhislysleuaspglycys 930935940 proleuargalaaspaspleuileprothrglygluglygluphephe 945950955960 valserpropheseralaglugluglyaspphehisglnilehisala 965970975 aspleuasnalaalaglnasnleuglnargargleutrpseraspphe 980985990 aspileserglnileargleuargcysasptrpglygluvalaspgly 99510001005 gluprovalleuileproargthrthrglylysargthralaasp 101010151020 sertyrglyasnlysvalphetyrthrlysthrglyvalthrtyr 102510301035 tyrgluarggluargglylyslysargarglysvalphealagln 104010451050 glugluleuserglugluglualagluleuleuvalglualaasp 105510601065 glualaargglulysservalvalleumetargaspprosergly 107010751080 ileileasnargglyasptrpthrargglnlysgluphetrpser 108510901095 metvalasnglnargilegluglytyrleuvallysglnilearg 110011051110 serargvalargleuglngluseralacysgluasnthrglyasp 111511201125 ile 2 1147 prt alicyclobacilluskakegawensis 2 metalavallysserilelysvallysleuargleuserglucyspro 151015 aspileleualaglymettrpglnleuhisargalathrasnalagly 202530 valargtyrtyrthrglutrpvalserleumetargglngluileleu 354045 tyrserargglyproaspglyglyglnglncystyrmetthralaglu 505560 aspcysglnarggluleuleuargargleuargasnargglnleuhis 65707580 asnglyargglnaspglnproglythraspalaaspleuleualaile 859095 serargargleutyrgluileleuvalleuglnserileglylysarg 100105110 glyaspalaglnglnilealaserserpheleuserproleuvalasp 115120125 proasnserlysglyglyargglyglualalysserglyarglyspro 130135140 alatrpglnlysmetargaspglnglyaspproargtrpvalalaala 145150155160 argglulystyrgluglnarglysalavalaspproserlysgluile 165170175 leuasnserleuaspalaleuglyleuargproleuphealavalphe 180185190 thrgluthrtyrargserglyvalasptrplysproleuglylysser 195200205 glnglyvalargthrtrpaspargaspmetpheglnglnalaleuglu 210215220 argleumetsertrpglusertrpasnargargvalglygluglutyr 225230235240 alaargleupheglnglnlysmetlysphegluglngluhispheala 245250255 gluglnserhisleuvallysleualaargalaleuglualaaspmet 260265270 argalaalaserglnglypheglualalysargglythralahisgln 275280285 ilethrargargalaleuargglyalaaspargvalphegluiletrp 290295300 lysserileprogluglualaleupheserglntyraspgluvalile 305310315320 argglnvalglnalaglulysargargasppheglyserhisaspleu 325330335 phealalysleualagluprolystyrglnproleutrpargalaasp 340345350 gluthrpheleuthrargtyralaleutyrasnglyvalleuargasp 355360365 leuglulysalaargglnphealathrphethrleuproaspalacys 370375380 valasnproiletrpthrargphegluserserglnglyserasnleu 385390395400 hislystyrglupheleupheasphisleuglyproglyarghisala 405410415 valargpheglnargleuleuvalvalglusergluglyalalysglu 420425430 argaspservalvalvalprovalalaproserglyglnleuasplys 435440445 leuvalleuargglugluglulysserservalalaleuhisleuhis 450455460 aspthralaargproaspglyphemetalaglutrpalaglyalalys 465470475480 leuglntyrgluargserthrleualaarglysalaargargasplys 485490495 glnglymetargsertrpargargglnprosermetleumetserala 500505510 alaglnmetleugluaspalalysglnalaglyaspvaltyrleuasn 515520525 ileservalargvallysserprosergluvalargglyglnargarg 530535540 proprotyralaalaleupheargileaspasplysglnargargval 545550555560 thrvalasntyrasnlysleuseralatyrleuglugluhisproasp 565570575 lysglnileproglyalaproglyleuleuserglyleuargvalmet 580585590 servalaspleuglyleuargthrseralaserileservalphearg 595600605 valalalyslysglugluvalglualaleuglyaspglyargpropro 610615620 histyrtyrproilehisglythraspaspleuvalalavalhisglu 625630635640 argserhisleuileglnmetproglygluthrgluthrlysglnleu 645650655 arglysleuargglugluargglnalavalleuargproleupheala 660665670 glnleualaleuleuargleuleuvalargcysglyalaalaaspglu 675680685 argileargthrargsertrpglnargleuthrlysglnglyargglu 690695700 phethrlysargleuthrprosertrpargglualaleugluleuglu 705710715720 leuthrargleuglualatyrcysglyargvalproaspaspglutrp 725730735 serargilevalaspargthrvalilealaleutrpargargmetgly 740745750 lysglnvalargasptrparglysglnvallysserglyalalysval 755760765 lysvallysglytyrglnleuaspvalvalglyglyasnserleuala 770775780 glnileasptyrleugluglnglntyrlyspheleuargargtrpser 785790795800 phephealaargalaserglyleuvalvalargalaasparggluser 805810815 hisphealavalalaleuargglnhisilegluasnalalysargasp 820825830 argleulyslysleualaaspargileleumetglualaleuglytyr 835840845 valtyrglualaserglyproarggluglyglntrpthralaglnhis 850855860 proprocysglnleuileileleuglugluleuseralatyrargphe 865870875880 seraspaspargproprosergluasnserlysleumetalatrpgly 885890895 hisargglyileleuglugluleuvalasnglnalaglnvalhisasp 900905910 valleuvalglythrvaltyralaalapheserserargpheaspala 915920925 argthrglyalaproglyvalargcysargargvalproalaargphe 930935940 valglyalathrvalaspaspserleuproleutrpleuthrgluphe 945950955960 leuasplyshisargleuasplysasnleuleuargproaspaspval 965970975 ileprothrglygluglyglupheleuvalserprocysglygluglu 980985990 alaalaargvalargglnvalhisalaaspileasnalaalaglnasn 99510001005 leuglnargargleutrpglnasnpheaspilethrgluleuarg 101010151020 leuargcysaspvallysmetglyglygluglythrvalleuval 102510301035 proargvalasnasnalaargalalysglnleupheglylyslys 104010451050 valleuvalserglnaspglyvalthrphephegluargsergln 105510601065 thrglyglylysprohisserglulysglnthraspleuthrasp 107010751080 lysgluleugluleuilealaglualaaspglualaargalalys 108510901095 servalvalleupheargaspproserglyhisileglylysgly 110011051110 histrpileargglnarggluphetrpserleuvallysglnarg 111511201125 ilegluserhisthralagluargileargvalargglyvalgly 113011351140 serserleuasp 1145 3 1146 prt alicyclobacillusmacrosporangiidus 3 metasnvalalavallysserilelysvallysleumetleuglyhis 151015 leuprogluilearggluglyleutrphisleuhisglualavalasn 202530 leuglyvalargtyrtyrthrglutrpleualaleuleuargglngly 354045 asnleutyrargargglylysaspglyalaglnglucystyrmetthr 505560 alagluglncysargglngluleuleuvalargleuargasparggln 65707580 lysargasnglyhisthrglyaspproglythraspglugluleuleu 859095 glyvalalaargargleutyrgluleuleuvalproglnservalgly 100105110 lyslysglyglnalaglnmetleualaserglypheleuserproleu 115120125 alaaspprolyssergluglyglylysglythrserlysserglyarg 130135140 lysproalatrpmetglymetlysglualaglyaspserargtrpval 145150155160 glualalysalaargtyrglualaasnlysalalysaspprothrlys 165170175 glnvalilealaserleuglumettyrglyleuargproleupheasp 180185190 valphethrgluthrtyrlysthrileargtrpmetproleuglylys 195200205 hisglnglyvalargalatrpaspargaspmetpheglnglnserleu 210215220 gluargleumetsertrpglusertrpasngluargvalglyalaglu 225230235240 phealaargleuvalaspargargaspargpheargglulyshisphe 245250255 thrglyglngluhisleuvalalaleualaglnargleugluglnglu 260265270 metlysglualaserproglyphegluserlysserserglnalahis 275280285 argilethrlysargalaleuargglyalaaspglyileileaspasp 290295300 trpleulysleusergluglygluprovalaspargpheaspgluile 305310315320 leuarglysargglnalaglnasnproargargpheglyserhisasp 325330335 leupheleulysleualagluprovalpheglnproleutrpargglu 340345350 aspproserpheleuserargtrpalasertyrasngluvalleuasn 355360365 lysleugluaspalalysglnphealathrphethrleuproserpro 370375380 cysserasnprovaltrpalaargphegluasnalagluglythrasn 385390395400 ilephelystyrasppheleupheasphispheglylysglyarghis 405410415 glyvalargpheglnargmetilevalmetargaspglyvalprothr 420425430 gluvalgluglyilevalvalproilealaproserargglnleuasp 435440445 alaleualaproasnaspalaalaserproileaspvalphevalgly 450455460 aspproalaalaproglyalapheargglyglnpheglyglyalalys 465470475480 ileglntyrargargseralaleuvalarglysglyargarggluglu 485490495 lysalatyrleucysglypheargleuproserglnargargthrgly 500505510 thrproalaaspaspalaglygluvalpheleuasnleuserleuarg 515520525 valgluserglnsergluglnalaglyargargasnproprotyrala 530535540 alavalphehisileseraspglnthrargargvalilevalargtyr 545550555560 glygluilegluargtyrleualagluhisproaspthrglyilepro 565570575 glyserargglyleuthrserglyleuargvalmetservalaspleu 580585590 glyleuargthrseralaalaileservalpheargvalalahisarg 595600605 aspgluleuthrproaspalahisglyargglnprophephephepro 610615620 ilehisglymetasphisleuvalalaleuhisgluargserhisleu 625630635640 ileargleuproglygluthrgluserlyslysvalargserilearg 645650655 gluglnargleuaspargleuasnargleuargserglnmetalaser 660665670 leuargleuleuvalargthrglyvalleuaspgluglnlysargasp 675680685 argasntrpgluargleuglnsersermetgluargglyglygluarg 690695700 metproserasptrptrpaspleupheglnalaglnvalargtyrleu 705710715720 alaglnhisargaspalaserglyglualatrpglyargmetvalgln 725730735 alaalavalargthrleutrpargglnleualalysglnvalargasp 740745750 trparglysgluvalargargasnalaasplysvallysilearggly 755760765 ilealaargaspvalproglyglyhisserleualaglnleuasptyr 770775780 leugluargglntyrargpheleuargsertrpseralapheserval 785790795800 glnalaglyglnvalvalargalagluargaspserargphealaval 805810815 alaleuarggluhisileaspasnglylyslysaspargleulyslys 820825830 leualaaspargileleumetglualaleuglytyrvaltyrvalthr 835840845 aspglyargargalaglyglntrpglnalavaltyrproprocysgln 850855860 leuvalleuleuglugluleuserglutyrargpheserasnasparg 865870875880 proprosergluasnserglnleumetvaltrpserhisargglyval 885890895 leuglugluleuilehisglnalaglnvalhisaspvalleuvalgly 900905910 thrileproalaalapheserserargpheaspalaargthrglyala 915920925 proglyileargcysargargvalproserileproleulysaspala 930935940 proserileproiletrpleuserhistyrleulysglnthrgluarg 945950955960 aspalaalaalaleuargproglygluleuileprothrglyaspgly 965970975 glupheleuvalthrproalaglyargglyalaserglyvalargval 980985990 valhisalaaspileasnalaalahisasnleuglnargargleutrp 99510001005 gluasnpheaspleuseraspileargvalargcysaspargarg 101010151020 gluglylysaspglythrvalvalleuileproargleuthrasn 102510301035 glnargvallysgluargtyrserglyvalilephethrserglu 104010451050 aspglyvalserphethrvalglyaspalalysthrargargarg 105510601065 serseralaserglnglygluglyaspaspleuseraspgluglu 107010751080 glngluleuleualaglualaaspaspalaarggluargserval 108510901095 valleupheargaspproserglyphevalasnglyglyargtrp 110011051110 thralaglnargalaphetrpglymetvalhisasnargileglu 111511201125 thrleuleualagluargpheservalserglyalaalaglulys 113011351140 valarggly 1145 4 1108 prt bacillushisashii 4 metalathrargserpheileleulysilegluproasnglugluval 151015 lyslysglyleutrplysthrhisgluvalleuasnhisglyileala 202530 tyrtyrmetasnileleulysleuileargglnglualailetyrglu 354045 hishisgluglnaspprolysasnprolyslysvalserlysalaglu 505560 ileglnalagluleutrpaspphevalleulysmetglnlyscysasn 65707580 serphethrhisgluvalasplysaspgluvalpheasnileleuarg 859095 gluleutyrglugluleuvalproserservalglulyslysglyglu 100105110 alaasnglnleuserasnlyspheleutyrproleuvalaspproasn 115120125 serglnserglylysglythralaserserglyarglysproargtrp 130135140 tyrasnleulysilealaglyaspprosertrpglugluglulyslys 145150155160 lystrpglugluasplyslyslysaspproleualalysileleugly 165170175 lysleualaglutyrglyleuileproleupheileprotyrthrasp 180185190 serasngluproilevallysgluilelystrpmetglulysserarg 195200205 asnglnservalargargleuasplysaspmetpheileglnalaleu 210215220 gluargpheleusertrpglusertrpasnleulysvallysgluglu 225230235240 tyrglulysvalglulysglutyrlysthrleuglugluargilelys 245250255 gluaspileglnalaleulysalaleugluglntyrglulysgluarg 260265270 glngluglnleuleuargaspthrleuasnthrasnglutyrargleu 275280285 serlysargglyleuargglytrparggluileileglnlystrpleu 290295300 lysmetaspgluasngluproserglulystyrleugluvalphelys 305310315320 asptyrglnarglyshisproargglualaglyasptyrservaltyr 325330335 glupheleuserlyslysgluasnhispheiletrpargasnhispro 340345350 glutyrprotyrleutyralathrphecysgluileasplyslyslys 355360365 lysaspalalysglnglnalathrphethrleualaaspproileasn 370375380 hisproleutrpvalargpheglugluargserglyserasnleuasn 385390395400 lystyrargileleuthrgluglnleuhisthrglulysleulyslys 405410415 lysleuthrvalglnleuaspargleuiletyrprothrglusergly 420425430 glytrpgluglulysglylysvalaspilevalleuleuproserarg 435440445 glnphetyrasnglnilepheleuaspilegluglulysglylyshis 450455460 alaphethrtyrlysaspgluserilelyspheproleulysglythr 465470475480 leuglyglyalaargvalglnpheaspargasphisleuargargtyr 485490495 prohislysvalgluserglyasnvalglyargiletyrpheasnmet 500505510 thrvalasnilegluprothrgluserprovalserlysserleulys 515520525 ilehisargaspasppheprolysvalvalasnphelysprolysglu 530535540 leuthrglutrpilelysaspserlysglylyslysleulyssergly 545550555560 ilegluserleugluileglyleuargvalmetserileaspleugly 565570575 glnargglnalaalaalaalaserilephegluvalvalaspglnlys 580585590 proaspilegluglylysleuphepheproilelysglythrgluleu 595600605 tyralavalhisargalaserpheasnilelysleuproglygluthr 610615620 leuvallysserarggluvalleuarglysalaarggluaspasnleu 625630635640 lysleumetasnglnlysleuasnpheleuargasnvalleuhisphe 645650655 glnglnphegluaspilethrgluargglulysargvalthrlystrp 660665670 ileserargglngluasnseraspvalproleuvaltyrglnaspglu 675680685 leuileglnilearggluleumettyrlysprotyrlysasptrpval 690695700 alapheleulysglnleuhislysargleugluvalgluileglylys 705710715720 gluvallyshistrparglysserleuseraspglyarglysglyleu 725730735 tyrglyileserleulysasnileaspgluileaspargthrarglys 740745750 pheleuleuargtrpserleuargprothrgluproglygluvalarg 755760765 argleugluproglyglnargphealaileaspglnleuasnhisleu 770775780 asnalaleulysgluaspargleulyslysmetalaasnthrileile 785790795800 methisalaleuglytyrcystyraspvalarglyslyslystrpgln 805810815 alalysasnproalacysglnileileleuphegluaspleuserasn 820825830 tyrasnprotyrglugluargserargphegluasnserlysleumet 835840845 lystrpserargarggluileproargglnvalalaleuglnglyglu 850855860 iletyrglyleuglnvalglygluvalglyalaglnpheserserarg 865870875880 phehisalalysthrglyserproglyileargcysservalvalthr 885890895 lysglulysleuglnaspasnargphephelysasnleuglnargglu 900905910 glyargleuthrleuasplysilealavalleulysgluglyaspleu 915920925 tyrproasplysglyglyglulyspheileserleuserlysasparg 930935940 lyscysvalthrthrhisalaaspileasnalaalaglnasnleugln 945950955960 lysargphetrpthrargthrhisglyphetyrlysvaltyrcyslys 965970975 alatyrglnvalaspglyglnthrvaltyrileprogluserlysasp 980985990 glnlysglnlysileilegluglupheglygluglytyrpheileleu 99510001005 lysaspglyvaltyrglutrpvalasnalaglylysleulysile 101010151020 lyslysglyserserlysglnsersersergluleuvalaspser 102510301035 aspileleulysaspserpheaspleualasergluleulysgly 104010451050 glulysleumetleutyrargaspproserglyasnvalphepro 105510601065 serasplystrpmetalaalaglyvalphepheglylysleuglu 107010751080 argileleuileserlysleuthrasnglntyrserileserthr 108510901095 ilegluaspaspserserlysglnsermet 11001105 5 1108 prt bacillus 5 metalaileargserilelysleulysleulysthrhisthrglypro 151015 glualaglnasnleuarglysglyiletrpargthrhisargleuleu 202530 asngluglyvalalatyrtyrmetlysmetleuleuleuphearggln 354045 gluserthrglygluargprolysglugluleuglnglugluleuile 505560 cyshisilearggluglnglnglnargasnglnalaasplysasnthr 65707580 glnalaleuproleuasplysalaleuglualaleuargglnleutyr 859095 gluleuleuvalproserservalglyglnserglyaspalaglnile 100105110 ileserarglyspheleuserproleuvalaspproasnserglugly 115120125 glylysglythrserlysalaglyalalysprothrtrpglnlyslys 130135140 lysglualaasnaspprothrtrpgluglnasptyrglulystrplys 145150155160 lysargargglugluaspprothralaservalilethrthrleuglu 165170175 glutyrglyileargproilepheproleutyrthrasnthrvalthr 180185190 aspilealatrpleuproleuglnserasnglnphevalargthrtrp 195200205 aspargaspmetleuglnglnalailegluargleuleusertrpglu 210215220 sertrpasnlysargvalglngluglutyralalysleulysglulys 225230235240 metalaglnleuasngluglnleugluglyglyglnglutrpileser 245250255 leuleugluglntyrglugluasnarggluarggluleuarggluasn 260265270 metthralaalaasnasplystyrargilethrlysargglnmetlys 275280285 glytrpasngluleutyrgluleutrpserthrpheproalaserala 290295300 serhisgluglntyrlysglualaleulysargvalglnglnargleu 305310315320 argglyargpheglyaspalahisphepheglntyrleumetgluglu 325330335 lysasnargleuiletrplysglyasnproglnargilehistyrphe 340345350 valalaargasngluleuthrlysargleugluglualalysglnser 355360365 alathrmetthrleuproasnalaarglyshisproleutrpvalarg 370375380 pheaspalaargglyglyasnleuglnasptyrtyrleuthralaglu 385390395400 alaasplysproargserargargphevalthrpheserglnleuile 405410415 trpprosergluserglytrpmetglulyslysaspvalgluvalglu 420425430 leualaleuserargglnphetyrglnglnvallysleuleulysasn 435440445 asplysglylysglnlysilegluphelysasplysglyserglyser 450455460 thrpheasnglyhisleuglyglyalalysleuglnleugluarggly 465470475480 aspleuglulysgluglulysasnphegluaspglygluileglyser 485490495 valtyrleuasnvalvalileaspphegluproleuglngluvallys 500505510 asnglyargvalglnalaprotyrglyglnvalleuglnleuilearg 515520525 argproasnglupheprolysvalthrthrtyrlyssergluglnleu 530535540 valglutrpilelysalaserproglnhisseralaglyvalgluser 545550555560 leualaserglypheargvalmetserileaspleuglyleuargala 565570575 alaalaalathrserilepheservalglugluserserasplysasn 580585590 alaalaaspphesertyrtrpilegluglythrproleuvalalaval 595600605 hisglnargsertyrmetleuargleuproglygluglnvalglulys 610615620 glnvalmetglulysargaspgluargpheglnleuhisglnargval 625630635640 lyspheglnileargvalleualaglnilemetargmetalaasnlys 645650655 glntyrglyaspargtrpaspgluleuaspserleulysglnalaval 660665670 gluglnlyslysserproleuaspglnthraspargthrphetrpglu 675680685 glyilevalcysaspleuthrlysvalleuproargasnglualaasp 690695700 trpgluglnalavalvalglnilehisarglysalagluglutyrval 705710715720 glylysalavalglnalatrparglysargphealaalaaspgluarg 725730735 lysglyilealaglyleusermettrpasnileglugluleuglugly 740745750 leuarglysleuleuilesertrpserargargthrargasnprogln 755760765 gluvalasnargphegluargglyhisthrserhisglnargleuleu 770775780 thrhisileglnasnvallysgluaspargleulysglnleuserhis 785790795800 alailevalmetthralaleuglytyrvaltyraspgluarglysgln 805810815 glutrpcysalaglutyrproalacysglnvalileleuphegluasn 820825830 leuserglntyrargserasnleuaspargserthrlysgluasnser 835840845 thrleumetlystrpalahisargserileprolystyrvalhismet 850855860 glnalagluprotyrglyileglnileglyaspvalargalaglutyr 865870875880 serserargphetyralalysthrglythrproglyileargcyslys 885890895 lysvalargglyglnaspleuglnglyargargphegluasnleugln 900905910 lysargleuvalasngluglnpheleuthrglugluglnvallysgln 915920925 leuargproglyaspilevalproaspaspserglygluleuphemet 930935940 thrleuthraspglyserglyserlysgluvalvalpheleuglnala 945950955960 aspileasnalaalahisasnleuglnlysargphetrpglnargtyr 965970975 asngluleuphelysvalsercysargvalilevalargaspgluglu 980985990 glutyrleuvalprolysthrlysservalglnalalysleuglylys 99510001005 glyleuphevallyslysseraspthralatrplysaspvaltyr 101010151020 valtrpaspserglnalalysleulysglylysthrthrphethr 102510301035 gluglusergluserprogluglnleugluasppheglngluile 104010451050 ilegluglualagluglualalysglythrtyrargthrleuphe 105510601065 argaspproserglyvalphepheprogluservaltrptyrpro 107010751080 glnlysaspphetrpglygluvallysarglysleutyrglylys 108510901095 leuarggluargpheleuthrlysalaarg 11001105 6 1112 prt bacillus 6 metalaileargserilelysleulysmetlysthrasnserglythr 151015 aspseriletyrleuarglysalaleutrpargthrhisglnleuile 202530 asngluglyilealatyrtyrmetasnleuleuthrleutyrarggln 354045 glualaileglyasplysthrlysglualatyrglnalagluleuile 505560 asnileileargasnglnglnargasnasnglyserserglugluhis 65707580 glyseraspglngluileleualaleuleuargglnleutyrgluleu 859095 ileileproserserileglygluserglyaspalaasnglnleugly 100105110 asnlyspheleutyrproleuvalaspproasnserglnserglylys 115120125 glythrserasnalaglyarglysproargtrplysargleulysglu 130135140 gluglyasnproasptrpgluleuglulyslyslysaspglugluarg 145150155160 lysalalysaspprothrvallysilepheaspasnleuasnlystyr 165170175 glyleuleuproleupheproleuphethrasnileglnlysaspile 180185190 glutrpleuproleuglylysargglnservalarglystrpasplys 195200205 aspmetpheileglnalailegluargleuleusertrpglusertrp 210215220 asnargargvalalaaspglutyrlysglnleulysglulysthrglu 225230235240 sertyrtyrlysgluhisleuthrglyglygluglutrpileglulys 245250255 ilearglyspheglulysgluargasnmetgluleuglulysasnala 260265270 phealaproasnaspglytyrpheilethrserargglnilearggly 275280285 trpaspargvaltyrglulystrpserlysleuprogluseralaser 290295300 proglugluleutrplysvalvalalagluglnglnasnlysmetser 305310315320 gluglypheglyaspprolysvalpheserpheleualaasnargglu 325330335 asnargaspiletrpargglyhissergluargiletyrhisileala 340345350 alatyrasnglyleuglnlyslysleuserargthrlysgluglnala 355360365 thrphethrleuproaspalailegluhisproleutrpileargtyr 370375380 gluserproglyglythrasnleuasnleuphelysleugluglulys 385390395400 glnlyslysasntyrtyrvalthrleuserlysileiletrpproser 405410415 gluglulystrpileglulysgluasnilegluileproleualapro 420425430 serileglnpheasnargglnilelysleulysglnhisvallysgly 435440445 lysglngluileserpheserasptyrserserargileserleuasp 450455460 glyvalleuglyglyserargileglnpheasnarglystyrilelys 465470475480 asnhislysgluleuleuglygluglyaspileglyprovalphephe 485490495 asnleuvalvalaspvalalaproleuglngluthrargasnglyarg 500505510 leuglnserproileglylysalaleulysvalileserseraspphe 515520525 serlysvalileasptyrlysprolysgluleumetasptrpmetasn 530535540 thrglyseralaserasnserpheglyvalalaserleuleuglugly 545550555560 metargvalmetserileaspmetglyglnargthrseralaserval 565570575 serilephegluvalvallysgluleuprolysaspglngluglnlys 580585590 leuphetyrserileasnaspthrgluleuphealailehislysarg 595600605 serpheleuleuasnleuproglygluvalvalthrlysasnasnlys 610615620 glnglnargglngluargarglyslysargglnphevalargsergln 625630635640 ileargmetleualaasnvalleuargleugluthrlyslysthrpro 645650655 aspgluarglyslysalailehislysleumetgluilevalglnser 660665670 tyraspsertrpthralaserglnlysgluvaltrpglulysgluleu 675680685 asnleuleuthrasnmetalaalapheasnaspgluiletrplysglu 690695700 serleuvalgluleuhishisargilegluprotyrvalglyglnile 705710715720 valserlystrparglysglyleusergluglyarglysasnleuala 725730735 glyilesermettrpasnileaspgluleugluaspthrargargleu 740745750 leuilesertrpserlysargserargthrproglyglualaasnarg 755760765 ilegluthraspglupropheglyserserleuleuglnhisilegln 770775780 asnvallysaspaspargleulysglnmetalaasnleuileilemet 785790795800 thralaleuglyphelystyrasplysgluglulysaspargtyrlys 805810815 argtrplysgluthrtyrproalacysglnileileleuphegluasn 820825830 leuasnargtyrleupheasnleuaspargserargarggluasnser 835840845 argleumetlystrpalahisargserileproargthrvalsermet 850855860 glnglyglumetpheglyleuglnvalglyaspvalargserglutyr 865870875880 serserargphehisalalysthrglyalaproglyileargcyshis 885890895 alaleuthrglugluaspleulysalaglyserasnthrleulysarg 900905910 leuilegluaspglypheileasnglusergluleualatyrleulys 915920925 lysglyaspileileproserglnglyglygluleuphevalthrleu 930935940 serlysargtyrlyslysaspseraspasnasngluleuthrvalile 945950955960 hisalaaspileasnalaalaglnasnleuglnlysargphetrpgln 965970975 glnasnsergluvaltyrargvalprocysglnleualaargmetgly 980985990 gluasplysleutyrileprolysserglnthrgluthrilelyslys 99510001005 tyrpheglylysglyserphevallysasnasnthrgluglnglu 101010151020 valtyrlystrpglulysserglulysmetlysilelysthrasp 102510301035 thrthrpheaspleuglnaspleuaspglyphegluaspileser 104010451050 lysthrilegluleualaglngluglnglnlyslystyrleuthr 105510601065 metpheargaspproserglytyrphepheasnasngluthrtrp 107010751080 argproglnlysglutyrtrpserilevalasnasnileilelys 108510901095 sercysleulyslyslysileleuserasnlysvalgluleu 110011051110 7 1149 prt desulfovibrioinopinatus 7 metprothrargthrileasnleulysleuvalleuglylysasnpro 151015 gluasnalathrleuargargalaleupheserthrhisargleuval 202530 asnglnalathrlysargilegluglupheleuleuleucysarggly 354045 glualatyrargthrvalaspasngluglylysglualagluilepro 505560 arghisalavalglngluglualaleualaphealalysalaalagln 65707580 arghisasnglycysileserthrtyrgluaspglngluileleuasp 859095 valleuargglnleutyrgluargleuvalproservalasngluasn 100105110 asnglualaglyaspalaglnalaalaasnalatrpvalserproleu 115120125 metseralaglusergluglyglyleuservaltyrasplysvalleu 130135140 aspproproprovaltrpmetlysleulysgluglulysalaprogly 145150155160 trpglualaalaserglniletrpileglnseraspgluglyglnser 165170175 leuleuasnlysproglyserproproargtrpilearglysleuarg 180185190 serglyglnprotrpglnaspaspphevalseraspglnlyslyslys 195200205 glnaspgluleuthrlysglyasnalaproleuilelysglnleulys 210215220 glumetglyleuleuproleuvalasnprophephearghisleuleu 225230235240 aspprogluglylysglyvalserprotrpaspargleualavalarg 245250255 alaalavalalahispheilesertrpglusertrpasnhisargthr 260265270 argalaglutyrasnserleulysleuargargaspgluphegluala 275280285 alaseraspgluphelysaspaspphethrleuleuargglntyrglu 290295300 alalysarghisserthrleulysserilealaleualaaspaspser 305310315320 asnprotyrargileglyvalargserleuargalatrpasnargval 325330335 arggluglutrpileasplysglyalathrglugluglnargvalthr 340345350 ileleuserlysleuglnthrglnleuargglylyspheglyasppro 355360365 aspleupheasntrpleualaglnasparghisvalhisleutrpser 370375380 proargaspservalthrproleuvalargileasnalavalasplys 385390395400 valleuargargarglysprotyralaleumetthrphealahispro 405410415 argphehisproargtrpileleutyrglualaproglyglyserasn 420425430 leuargglntyralaleuaspcysthrgluasnalaleuhisilethr 435440445 leuproleuleuvalaspaspalahisglythrtrpileglulyslys 450455460 ileargvalproleualaproserglyglnileglnaspleuthrleu 465470475480 glulysleuglulyslyslysasnargleutyrtyrargserglyphe 485490495 glnglnphealaglyleualaglyglyalagluvalleuphehisarg 500505510 protyrmetgluhisaspgluargserglugluserleuleugluarg 515520525 proglyalavaltrpphelysleuthrleuaspvalalathrglnala 530535540 proproasntrpleuaspglylysglyargvalargthrproproglu 545550555560 valhishisphelysthralaleuserasnlysserlyshisthrarg 565570575 thrleuglnproglyleuargvalleuservalaspleuglymetarg 580585590 thrphealasercysservalphegluleuilegluglylysproglu 595600605 thrglyargalapheprovalalaaspgluargsermetaspserpro 610615620 asnlysleutrpalalyshisgluargserphelysleuthrleupro 625630635640 glygluthrproserarglysglugluglugluargserilealaarg 645650655 alagluiletyralaleulysargaspileglnargleulysserleu 660665670 leuargleuglyglugluaspasnaspasnargargaspalaleuleu 675680685 gluglnphephelysglytrpglyglugluaspvalvalproglygln 690695700 alapheproargserleupheglnglyleuglyalaalaprophearg 705710715720 serthrprogluleutrpargglnhiscysglnthrtyrtyrasplys 725730735 alaglualacysleualalyshisileserasptrparglysargthr 740745750 argproargprothrserargglumettrptyrlysthrargsertyr 755760765 hisglyglylysseriletrpmetleuglutyrleuaspalavalarg 770775780 lysleuleuleusertrpserleuargglyargthrtyrglyalaile 785790795800 asnargglnaspthralaargpheglyserleualaserargleuleu 805810815 hishisileasnserleulysgluaspargilelysthrglyalaasp 820825830 serilevalglnalaalaargglytyrileproleuprohisglylys 835840845 glytrpgluglnargtyrgluprocysglnleuileleuphegluasp 850855860 leualaargtyrargpheargvalaspargproargarggluasnser 865870875880 glnleumetglntrpasnhisargalailevalalagluthrthrmet 885890895 glnalagluleutyrglyglnilevalgluasnthralaalaglyphe 900905910 serserargphehisalaalathrglyalaproglyvalargcysarg 915920925 pheleuleugluargasppheaspasnaspleuprolysprotyrleu 930935940 leuarggluleusertrpmetleuglyasnthrlysvalgluserglu 945950955960 gluglulysleuargleuleuserglulysileargproglyserleu 965970975 valprotrpaspglyglygluglnphealathrleuhisprolysarg 980985990 glnthrleucysvalilehisalaaspmetasnalaalaglnasnleu 99510001005 glnargargphepheglyargcysglyglualapheargleuval 101010151020 cysglnprohisglyaspaspvalleuargleualaserthrpro 102510301035 glyalaargleuleuglyalaleuglnglnleugluasnglygln 104010451050 glyalaphegluleuvalargaspmetglyserthrserglnmet 105510601065 asnargphevalmetlysserleuglylyslyslysilelyspro 107010751080 leuglnaspasnasnglyaspaspgluleugluaspvalleuser 108510901095 valleuproglugluaspaspthrglyargilethrvalphearg 110011051110 aspserserglyilephepheprocysasnvaltrpileproala 111511201125 lysglnphetrpproalavalargalametiletrplysvalmet 113011351140 alaserhisserleugly 1145 8 1090 prt laceyellasediminis 8 metserileargserphelysleulysilelysthrlysserglyval 151015 asnalaglugluleuargargglyleutrpargthrhisglnleuile 202530 asnaspglyilealatyrtyrmetasntrpleuvalleuleuarggln 354045 gluaspleupheileargasnglugluthrasngluileglulysarg 505560 serlysglugluileglnglygluleuleugluargvalhislysgln 65707580 glnglnargasnglntrpserglygluvalaspaspglnthrleuleu 859095 glnthrleuarghisleutyrglugluilevalproservalilegly 100105110 lysserglyasnalaserleulysalaargphepheleuglyproleu 115120125 valaspproasnasnlysthrthrlysaspvalserlysserglypro 130135140 thrprolystrplyslysmetlysaspalaglyaspproasntrpval 145150155160 glnglutyrglulystyrmetalagluargglnthrleuvalargleu 165170175 gluglumetglyleuileproleuphepromettyrthraspgluval 180185190 glyaspilehistrpleuproglnalaserglytyrthrargthrtrp 195200205 aspargaspmetpheglnglnalailegluargleuleusertrpglu 210215220 sertrpasnargargvalarggluargargalaglnpheglulyslys 225230235240 thrhisaspphealaserargphesergluseraspvalglntrpmet 245250255 asnlysleuargglutyrglualaglnglnglulysserleugluglu 260265270 asnalaphealaproasngluprotyralaleuthrlyslysalaleu 275280285 argglytrpgluargvaltyrhissertrpmetargleuaspserala 290295300 alasergluglualatyrtrpglngluvalalathrcysglnthrala 305310315320 metargglyglupheglyaspproalailetyrglnpheleualagln 325330335 lysgluasnhisaspiletrpargglytyrprogluargvalileasp 340345350 phealagluleuasnhisleuglnarggluleuargargalalysglu 355360365 aspalathrphethrleuproaspservalasphisproleutrpval 370375380 argtyrglualaproglyglythrasnilehisglytyraspleuval 385390395400 glnaspthrlysargasnleuthrleuileleuasplyspheileleu 405410415 proaspgluasnglysertrphisgluvallyslysvalpropheser 420425430 leualalysserlysglnphehisargglnvaltrpleuglngluglu 435440445 glnlysglnlyslysarggluvalvalphetyrasptyrserthrasn 450455460 leuprohisleuglythrleualaglyalalysleuglntrpasparg 465470475480 asnpheleuasnlysargthrglnglnglnileglugluthrglyglu 485490495 ileglylysvalphepheasnileservalaspvalargproalaval 500505510 gluvallysasnglyargleuglnasnglyleuglylysalaleuthr 515520525 valleuthrhisproaspglythrlysilevalthrglytrplysala 530535540 gluglnleuglulystrpvalglygluserglyargvalserserleu 545550555560 glyleuaspserleusergluglyleuargvalmetserileaspleu 565570575 glyglnargthrseralathrvalservalphegluilethrlysglu 580585590 alaproaspasnprotyrlysphephetyrglnleugluglythrglu 595600605 leuphealavalhisglnargserpheleuleualaleuproglyglu 610615620 asnproproglnlysilelysglnmetarggluileargtrplysglu 625630635640 argasnargilelysglnglnvalaspglnleuseralaileleuarg 645650655 leuhislyslysvalasngluaspgluargileglnalaileasplys 660665670 leuleuglnlysvalalasertrpglnleuasnglugluilealathr 675680685 alatrpasnglnalaleuserglnleutyrserlysalalysgluasn 690695700 aspleuglntrpasnglnalailelysasnalahishisglnleuglu 705710715720 provalvalglylysglnileserleutrparglysaspleuserthr 725730735 glyargglnglyilealaglyleuserleutrpserileglugluleu 740745750 glualathrlyslysleuleuthrargtrpserlysargserargglu 755760765 proglyvalvallysargilegluargphegluthrphealalysgln 770775780 ileglnhishisileasnglnvallysgluasnargleulysglnleu 785790795800 alaasnleuilevalmetthralaleuglytyrlystyraspglnglu 805810815 glnlyslystrpilegluvaltyrproalacysglnvalvalleuphe 820825830 gluasnleuargsertyrargphesertyrgluargserargargglu 835840845 asnlyslysleumetglutrpserhisargserileprolysleuval 850855860 glnmetglnglygluleupheglyleuglnvalalaaspvaltyrala 865870875880 alatyrserserargtyrhisglyargthrglyalaproglyilearg 885890895 cyshisalaleuthrglualaaspleuargasngluthrasnileile 900905910 hisgluleuileglualaglypheilelysglugluhisargprotyr 915920925 leuglnglnglyaspleuvalprotrpserglyglygluleupheala 930935940 thrleuglnlysprotyraspasnproargileleuthrleuhisala 945950955960 aspileasnalaalaglnasnileglnlysargphetrphisproser 965970975 mettrppheargvalasncysgluservalmetgluglygluileval 980985990 thrtyrvalprolysasnlysthrvalhislyslysglnglylysthr 99510001005 pheargphevallysvalgluglyseraspvaltyrglutrpala 101010151020 lystrpserlysasnargasnlysasnthrpheserserilethr 102510301035 gluarglysproprosersermetileleupheargaspproser 104010451050 glythrphephelysgluglnglutrpvalgluglnlysthrphe 105510601065 trpglylysvalglnsermetileglnalatyrmetlyslysthr 107010751080 ilevalglnargmetgluglu 10851090 9 1119 prt spirochaetes 9 metserphethrilesertyrprophelysleuileilelysasnlys 151015 aspglualalysalaleuleuaspthrhisglntyrmetasnglugly 202530 vallystyrtyrleuglulysleuleumetpheargglnglulysile 354045 pheileglygluaspgluthrglylysargiletyrileglugluthr 505560 glutyrlyslysglnileglugluphetyrleuilelyslysthrglu 65707580 leuglyargasnleuthrleuthrleuaspgluphelysthrleumet 859095 arggluleutyrilecysleuvalsersersermetgluasnlyslys 100105110 glypheproasnalaglnglnalaserleuasnilepheserproleu 115120125 pheaspalagluserlysglytyrileleulysglugluasnasnasn 130135140 ileserleuilehislysasptyrglylysileleuleulysargleu 145150155160 argaspasnasnleuileproilephethrlysphethraspilelys 165170175 lysilethralalysleuserprothralaleuaspargmetilephe 180185190 alaglnalaileglulysleuleusertyrglusertrpcyslysleu 195200205 metilelysgluargpheasplysgluvallysilelysgluleuglu 210215220 asnlyscysgluasnlysglngluargasplysilephegluileleu 225230235240 glulystyrgluglugluargglnlysthrphegluglnaspsergly 245250255 phealalyslysglylysphetyrilethrglyargmetleulysgly 260265270 pheaspgluilelysglulystrpleulysglulysaspargserglu 275280285 glnasnleuileasnileleuasnlystyrglnthraspasnserlys 290295300 leuvalglyaspargasnleupheglupheileilelysleugluasn 305310315320 glncysleutrpasnglyaspileasptyrleulysilelysargasp 325330335 ileasnlysasnglniletrpleuaspargproglumetproargphe 340345350 thrmetproaspphelyslyshisproleutrptyrargtyrgluasp 355360365 proserasnserasnpheargasntyrlysilegluvalvallysasp 370375380 gluasntyrilethrileproleuilethrgluargasnasnglutyr 385390395400 pheglugluasntyrthrpheasnleualalysleulyslysleuser 405410415 gluasnilethrpheileprolysserlysasnlysgluphegluphe 420425430 ileaspserasnaspgluglugluasplyslysaspglnlyslysser 435440445 lysglntyrilelystyrcysaspthralalysasnthrsertyrgly 450455460 lysserglyglyileargleutyrpheasnargasngluleugluasn 465470475480 tyrlysaspglylyslysmetaspsertyrthrvalphethrleuser 485490495 ileargasptyrlysserleuphealalysglulysleuglnprogln 500505510 ilepheasnthrvalaspasnlysilethrserleulysileglnlys 515520525 lyspheglyasnglugluglnthrasnpheleusertyrphethrgln 530535540 asnglnilethrlyslysasptrpmetaspglulysthrpheglnasn 545550555560 vallysgluleuasngluglyileargvalleuservalaspleugly 565570575 glnargphephealaalavalsercysphegluilemetsergluile 580585590 aspasnasnlysleuphepheasnleuasnaspglnasnhislysile 595600605 ileargileasnasplysasntyrtyralalyshisiletyrserlys 610615620 thrilelysleuserglygluaspaspaspleutyrlysgluarglys 625630635640 ileasnlysasntyrlysleusertyrglngluarglysasnlysile 645650655 glyilephethrargglnileasnlysleuasnglnleuleulysile 660665670 ileargasnaspgluileasplysglulysphelysgluleuileglu 675680685 thrthrlysargtyrvallysasnthrtyrasnaspglyileileasp 690695700 trpasnasnvalaspasnlysileleusertyrgluasnlysgluasp 705710715720 valileasnleuhislysgluleuasplyslysleugluileaspphe 725730735 lysglupheileargglucysarglysproilepheargserglygly 740745750 leusermetglnargileasppheleuglulysleuasnlysleulys 755760765 arglystrpvalalaargthrglnlysseralagluserilevalleu 770775780 thrprolyspheglytyrlysleulysgluhisileasngluleulys 785790795800 aspasnargvallysglnglyvalasntyrileleumetthralaleu 805810815 glytyrilelysaspasngluilelysasnaspserlyslyslysgln 820825830 lysgluasptrpvallyslysasnargalacysglnileileleumet 835840845 glulysleuthrglutyrthrphealagluaspargproarggluglu 850855860 asnserlysleuargmettrpserhisargglnilepheasnpheleu 865870875880 glnglnlysalaserleutrpglyileleuvalglyaspvalpheala 885890895 protyrthrserlyscysleuseraspasnasnalaproglyilearg 900905910 cyshisglnvalthrlyslysaspleuileaspasnsertrppheleu 915920925 lysilevalvallysaspaspalaphecysaspleuilegluileasn 930935940 lysgluasnvallysasnlysserilelysileasnaspileleupro 945950955960 leuargglyglygluleuphealaserilelysaspglylysleuhis 965970975 ilevalglnalaaspileasnalaserargasnilealalysargphe 980985990 leuserglnileasnpropheargvalvalleulyslysasplysasp 99510001005 gluthrphehisleulysasngluproasntyrleulysasntyr 101010151020 tyrserileleuasnphevalprothrasnglugluleuthrphe 102510301035 phelysvalglugluasnlysaspilelysprothrlysargile 104010451050 lysmetasplyshisglulysgluserthraspgluglyaspasp 105510601065 tyrserlysasnglnilealaleupheargaspaspserglyile 107010751080 phepheasplysserleutrpvalaspglylysilephetrpser 108510901095 valvallysasnlysmetthrlysleuleuarggluargasnasn 110011051110 lyslysasnglyserlys 1115 10 1142 prt tuberibacilluscalidus 10 metasnilehisleulysgluleuileargmetalathrlysserphe 151015 ileleulysmetlysthrlysasnasnproglnleuargleuserleu 202530 trplysthrhisgluleupheasnpheglyvalalatyrtyrmetasp 354045 leuleuserleupheargglnlysaspleutyrmethisasnaspglu 505560 aspproasphisprovalvalleulyslysglugluileglngluarg 65707580 leutrpmetlysvalarggluthrglnglnlysasnglyphehisgly 859095 gluvalserlysaspgluvalleugluthrleuargalaleutyrglu 100105110 gluleuvalproseralavalglylysserglyglualaasnglnile 115120125 serasnlystyrleutyrproleuthraspproalaserglnsergly 130135140 lysglythralaasnserglyarglysproargtrplyslysleulys 145150155160 glualaglyaspprosertrplysaspalatyrglulystrpglulys 165170175 gluargglngluaspprolysleulysileleualaalaleuglnser 180185190 pheglyleuileproleupheargprophethrgluasnasphislys 195200205 alavalileservallystrpmetprolysserlysasnglnserval 210215220 arglyspheasplysaspmetpheasnglnalailegluargpheleu 225230235240 sertrpglusertrpasnglulysvalalagluasptyrglulysthr 245250255 valseriletyrgluserleuglnlysgluleulysglyileserthr 260265270 lysalaphegluilemetgluargvalglulysalatyrglualahis 275280285 leuarggluilethrpheserasnserthrtyrargileglyasnarg 290295300 alaileargglytrpthrgluilevallyslystrpmetlysleuasp 305310315320 proseralaproglnglyasntyrleuaspvalvallysasptyrgln 325330335 argarghisproarggluserglyaspphelysleuphegluleuleu 340345350 serargprogluasnglnalaalatrpargglutyrproglupheleu 355360365 proleutyrvallystyrarghisalagluglnargmetlysthrala 370375380 lyslysglnalathrphethrleucysaspproilearghisproleu 385390395400 trpvalargtyrglugluargserglythrasnleuasnlystyrarg 405410415 leuilemetasnglulysglulysvalvalglnpheaspargleuile 420425430 cysleuasnalaaspglyhistyrglugluglngluaspvalthrval 435440445 proleualaproserglnglnpheaspaspglnilelyspheserser 450455460 gluaspthrglylysglylyshisasnphesertyrtyrhislysgly 465470475480 ileasntyrgluleulysglythrleuglyglyalaargileglnphe 485490495 asparggluhisleuleuargargglnglyvallysalaglyasnval 500505510 glyargilepheleuasnvalthrleuasnilegluprometglnpro 515520525 pheserargserglyasnleuglnthrservalglylysalaleulys 530535540 valtyrvalaspglytyrprolysvalvalasnphelysprolysglu 545550555560 leuthrgluhisilelysgluserglulysasnthrleuthrleugly 565570575 valgluserleuprothrglyleuargvalmetservalaspleugly 580585590 glnargglnalaalaalaileserilephegluvalvalserglulys 595600605 proaspaspasnlysleuphetyrprovallysaspthraspleuphe 610615620 alavalhisargthrserpheasnilelysleuproglyglulysarg 625630635640 thrgluargargmetleugluglnglnlysargaspglnalailearg 645650655 aspleuserarglysleulyspheleulysasnvalleuasnmetgln 660665670 lysleuglulysthraspgluargglulysargvalasnargtrpile 675680685 lysasparggluarggluglugluasnprovaltyrvalglngluphe 690695700 glumetileserlysvalleutyrserprohisservaltrpvalasp 705710715720 glnleulysserilehisarglysleuglugluglnleuglylysglu 725730735 ileserlystrpargglnserileserglnglyargglnglyvaltyr 740745750 glyileserleulysasnilegluaspileglulysthrargargleu 755760765 leupheargtrpsermetargprogluasnproglygluvallysgln 770775780 leuglnproglygluargphealaileaspglnglnasnhisleuasn 785790795800 hisleulysaspaspargilelyslysleualaasnglnilevalmet 805810815 thralaleuglytyrargtyraspglylysarglyslystrpileala 820825830 lyshisproalacysglnleuvalleuphegluaspleuserargtyr 835840845 alaphetyraspgluargserargleugluasnargasnleumetarg 850855860 trpserargarggluileprolysglnvalalaglnileglyglyleu 865870875880 tyrglyleuleuvalglygluvalglyalaglntyrserserargphe 885890895 hisalalysserglyalaproglyileargcysargvalvallysglu 900905910 hisgluleutyrilethrgluglyglyglnlysvalargasnglnlys 915920925 pheleuaspserleuvalgluasnasnileilegluproaspaspala 930935940 argargleugluproglyaspleuileargaspglnglyglyasplys 945950955960 phealathrleuaspgluargglygluleuvalilethrhisalaasp 965970975 ileasnalaalaglnasnleuglnlysargphetrpthrargthrhis 980985990 glyleutyrargileargcysgluserarggluilelysaspalaval 99510001005 valleuvalproserasplysaspglnlysglulysmetgluasn 101010151020 leupheglyileglytyrleuglnprophelysglngluasnasp 102510301035 valtyrlystrpvallysglyglulysilelysglylyslysthr 104010451050 serserglnseraspasplysgluleuvalsergluileleugln 105510601065 glualaservalmetalaaspgluleulysglyasnarglysthr 107010751080 leupheargaspproserglytyrvalpheprolysaspargtrp 108510901095 tyrthrglyglyargtyrpheglythrleugluhisleuleulys 110011051110 arglysleualagluargargleupheaspglyglyserserarg 111511201125 argglyleupheasnglythraspserasnthrasnvalglu 113011351140 11 137 dna artificialsequence optimizedsgrnascaffold 11 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcgagc120 gatctgagaagtggcac137 12 141 dna artificialsequence optimizedsgrnascaffold 12 aactgtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaa60 agcccgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagc120 gagcgatctgagaagtggcac141 13 139 dna artificialsequence optimizedsgrnascaffold 13 ctgtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaag60 cccgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcga120 gcgatctgagaagtggcac139 14 127 dna artificialsequence optimizedsgrnascaffold 14 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaaaaagaacgctcgctcagtgttatcactgagcgagcgatctgagaa120 gtggcac127 15 99 dna artificialsequence optimizedsgrnascaffold 15 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaaaaagaacgatctgagaagtggcac99 16 93 dna artificialsequence optimizedsgrnascaffold 16 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaaaaagctgagaagtggcac93 17 91 dna artificialsequence optimizedsgrnascaffold 17 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaaagctgagaagtggcac91 18 91 dna artificialsequence optimizedsgrnascaffold 18 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaaaactgagaagtggcac91 19 89 dna artificialsequence optimizedsgrnascaffold 19 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaagcgagaagtggcac89 20 87 dna artificialsequence optimizedsgrnascaffold 20 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctaagcagaagtggcac87 21 85 dna artificialsequence optimizedsgrnascaffold 21 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttcaagcgaagtggcac85 22 137 dna artificialsequence aasgrna_scaffold 22 gtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagcc60 cgttgaacttctcaaaaagaacgctcgctcagtgttctgacgtcggatcactgagcgagc120 gatctgagaagtggcac137 23 137 dna artificialsequence aksgrna1_scaffold 23 tcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaaca60 ccccgttggcttcaagatgaccgctcgctcagcgatctgacaacggatcgctgagcgagc120 ggtctgagaagtggcac137 24 145 dna artificialsequence amsgrna1_scaffold 24 ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccagg60 agtgaacaccccgttggcttcaacatgatcgcccgctcaacggtccgatgtcggatcgtt120 gagcgggcgatctgagaagtggcac145 25 141 dna artificialsequence bhsgrna_scaffold 25 gaggttctgtcttttggtcaggacaaccgtctagctataagtgctgcagggtgtgagaaa60 ctcctattgctggacgatgtctcttttatttcttttttcttggatgtccaagaaaaaaga120 aatgatacgaggcattagcac141 26 132 dna artificialsequence bssgrna_scaffold 26 ccataagtcgacttacatatccgtgcgtgtgcattatgggcccatccacaggtctattcc60 cacggataatcacgactttccactaagctttcgaatgttcgaaagcttagtggaaagctt120 cgtggttagcac132 27 130 dna artificialsequence bs3sgrna_scaffold 27 ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccacc60 acaggattatcttatttctgctaagtgtttagttgcctgaatacttagcagaaataatga120 tgattggcac130 28 118 dna artificialsequence lssgrna_scaffold 28 ggcaaagaatactgtgcgtgtgctaaggatggaaaaaatccattcaaccacaggattaca60 ttatttatctaatcacttaaatctttaagtgattagatgaattaaatgtgattagcac118 29 126 dna artificialsequence sbsgrna_scaffold 29 gtcttagggtatatcccaaatttgtcttagtatgtgcattgcttacagcgacaactaagg60 tttgtttatcttttttttacattgtaagatgttttacattataaaaagaagataatctta120 ttgcac126 30 86 dna artificialsequence artsgrna1scaffold 30 ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagc60 ccgttgaacttcaagcgaagtggcac86 31 84 dna artificialsequence artsgrna2scaffold 31 ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccc60 cgttgacttcaagcgaagtggcac84 32 79 dna artificialsequence artsgrna3scaffold 32 ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacag60 acttcaagcgaagtggcac79 33 91 dna artificialsequence artsgrna4scaffold 33 ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggc60 aaagcccgttgaacttcaagcgaagtggcac91 34 89 dna artificialsequence artsgrna5scaffold 34 ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaa60 caccccgttgacttcaagcgaagtggcac89 35 84 dna artificialsequence artsgrna6scaffold 35 ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccac60 cacagacttcaagcgaagtggcac84 36 90 dna artificialsequence artsgrna7scaffold 36 ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggca60 aagcccgttgaacttcaagcgaagtggcac90 37 88 dna artificialsequence artsgrna8scaffold 37 ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaac60 accccgttgacttcaagcgaagtggcac88 38 83 dna artificialsequence artsgrna9scaffold 38 ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccacc60 acagacttcaagcgaagtggcac83 39 85 dna artificialsequence artsgrna10scaffold 39 ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagc60 ccgttgagcttcaaagaagtggcac85 40 83 dna artificialsequence artsgrna11scaffold 40 ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccc60 cgttggcttcaaagaagtggcac83 41 78 dna artificialsequence artsgrna12scaffold 41 ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacag60 gcttcaaagaagtggcac78 42 90 dna artificialsequence artsgrna13scaffold 42 ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggc60 aaagcccgttgagcttcaaagaagtggcac90 43 88 dna artificialsequence artsgrna14scaffold 43 ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaa60 caccccgttggcttcaaagaagtggcac88 44 83 dna artificialsequence artsgrna15scaffold 44 ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccac60 cacaggcttcaaagaagtggcac83 45 89 dna artificialsequence artsgrna16scaffold 45 ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggca60 aagcccgttgagcttcaaagaagtggcac89 46 87 dna artificialsequence artsgrna17scaffold 46 ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaac60 accccgttggcttcaaagaagtggcac87 47 82 dna artificialsequence artsgrna18scaffold 47 ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccacc60 acaggcttcaaagaagtggcac82 48 89 dna artificialsequence artsgrna19scaffold 48 ggtctaaaggacagaatttttcaacgggtgtgccaatggccactttccaggtggcaaagc60 ccgttgagattatctatgatgattggcac89 49 87 dna artificialsequence artsgrna20scaffold 49 ggtctaaaggacagaagacaacgggaagtgccaatgtgctctttccaagagcaaacaccc60 cgttggattatctatgatgattggcac87 50 82 dna artificialsequence artsgrna21scaffold 50 ggtctaaaggacagaaaatctgtgcgtgtgccataagtaattaaaaattacccaccacag60 gattatctatgatgattggcac82 51 94 dna artificialsequence artsgrna22scaffold 51 ggtcgtctataggacggcgagtttttcaacgggtgtgccaatggccactttccaggtggc60 aaagcccgttgagattatctatgatgattggcac94 52 92 dna artificialsequence artsgrna23scaffold 52 ggtcgtctataggacggcgaggacaacgggaagtgccaatgtgctctttccaagagcaaa60 caccccgttggattatctatgatgattggcac92 53 87 dna artificialsequence artsgrna24scaffold 53 ggtcgtctataggacggcgagaatctgtgcgtgtgccataagtaattaaaaattacccac60 cacaggattatctatgatgattggcac87 54 93 dna artificialsequence artsgrna25scaffold 54 ggtgacctatagggtcaatgtttttcaacgggtgtgccaatggccactttccaggtggca60 aagcccgttgagattatctatgatgattggcac93 55 91 dna artificialsequence artsgrna26scaffold 55 ggtgacctatagggtcaatggacaacgggaagtgccaatgtgctctttccaagagcaaac60 accccgttggattatctatgatgattggcac91 56 86 dna artificialsequence artsgrna27scaffold 56 ggtgacctatagggtcaatgaatctgtgcgtgtgccataagtaattaaaaattacccacc60 acaggattatctatgatgattggcac86 57 82 dna artificialsequence artsgrna28scaffold 57 ggtctaaaggacagaacaacgggatgtgccaatgcactctttccaggagtgaacaccccg60 ttgacttcaagcgaagtggcac82 58 87 dna artificialsequence artsgrna29scaffold 58 ggtcgtctataggacggcgagcaacgggatgtgccaatgcactctttccaggagtgaaca60 ccccgttgacttcaagcgaagtggcac87 59 99 dna artificialsequence artsgrna30scaffold 59 ggaattgccgatctataggacggcagatttttttcaacgggtgtgccaatggccactttc60 caggtggcaaagcccgttgaacttcaagcgaagtggcac99 60 97 dna artificialsequence artsgrna31scaffold 60 ggaattgccgatctataggacggcagattgacaacgggaagtgccaatgtgctctttcca60 agagcaaacaccccgttgacttcaagcgaagtggcac97 61 95 dna artificialsequence artsgrna32scaffold 61 ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccagg60 agtgaacaccccgttgacttcaagcgaagtggcac95 62 81 dna artificialsequence artsgrna33scaffold 62 ggtctaaaggacagaacaacgggatgtgccaatgcactctttccaggagtgaacaccccg60 ttggcttcaaagaagtggcac81 63 86 dna artificialsequence artsgrna34scaffold 63 ggtcgtctataggacggcgagcaacgggatgtgccaatgcactctttccaggagtgaaca60 ccccgttggcttcaaagaagtggcac86 64 98 dna artificialsequence artsgrna35scaffold 64 ggaattgccgatctataggacggcagatttttttcaacgggtgtgccaatggccactttc60 caggtggcaaagcccgttgagcttcaaagaagtggcac98 65 96 dna artificialsequence artsgrna36scaffold 65 ggaattgccgatctataggacggcagattgacaacgggaagtgccaatgtgctctttcca60 agagcaaacaccccgttggcttcaaagaagtggcac96 66 94 dna artificialsequence artsgrna37scaffold 66 ggaattgccgatctataggacggcagattcaacgggatgtgccaatgcactctttccagg60 agtgaacaccccgttggcttcaaagaagtggcac94 |
【本文地址】